Over 6,000 citations of NAMD reference paper
October 6, 2016
The NAMD developers thank our users for 6,000 citations of our 2005 reference paper:
James C. Phillips, Rosemary Braun, Wei Wang, James Gumbart, Emad Tajkhorshid, Elizabeth Villa, Christophe Chipot, Robert D. Skeel, Laxmikant Kale, and Klaus Schulten. Scalable molecular dynamics with NAMD. Journal of Computational Chemistry, 26:1781-1802, 2005.
NAMD is a parallel molecular dynamics code designed for high-performance simulation of large biomolecular systems. NAMD scales to hundreds of processors on high-end parallel platforms, as well as tens of processors on low-cost commodity clusters, and also runs on individual desktop and laptop computers. NAMD works with AMBER and CHARMM potential functions, parameters, and file formats. This article, directed to novices as well as experts, first introduces concepts and methods used in the NAMD program, describing the classical molecular dynamics force field, equations of motion, and integration methods along with the efficient electrostatics evaluation algorithms employed and temperature and pressure controls used. Features for steering the simulation across barriers and for calculating both alchemical and conformational free energy differences are presented. The motivations for and a roadmap to the internal design of NAMD, implemented in C++ and based on Charm++ parallel objects, are outlined. The factors affecting the serial and parallel performance of a simulation are discussed. Finally, typical NAMD use is illustrated with representative applications to a small, a medium, and a large biomolecular system, highlighting particular features of NAMD, for example, the Tcl scripting language. The article also provides a list of the key features of NAMD and discusses the benefits of combining NAMD with the molecular graphics/sequence analysis software VMD and the grid computing/collaboratory software BioCoRE. NAMD is distributed free of charge with source code at www.ks.uiuc.edu.
This paper is available from our website or directly from the journal.
The NAMD License Agreement requires that this paper be cited by any published work which utilizes NAMD. Proper citation is essential to continued NIH funding for NAMD development, as it is a primary way in which we demonstrate the value of our software to the scientific community. As of April 2016, over 7,000 publications have cited either the 1999 or 2005 reference papers.
Below are a few of the many biomedically, environmentally, and methodologically relevant discoveries made by NAMD users since our October 2015 milestone of 5,000 citations.
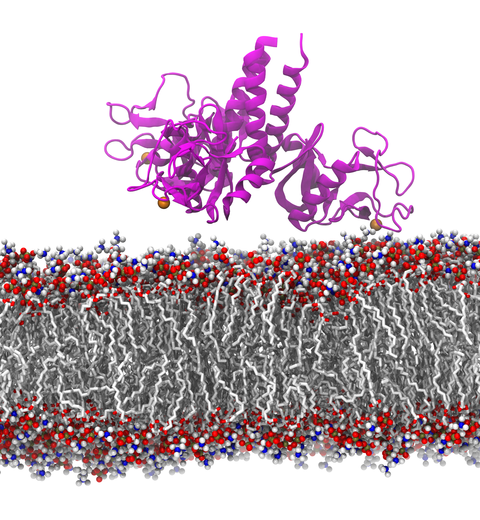
image size:
1.7MB
made with VMD
Our lungs are coated with a layer of protein and lipid mixture called lung surfactant, which prevents the lungs from collapsing and protects us from bacterial and viral infections (see October 2012 and January 2014 highlights). Lung surfactant protein A (SP-A) - the major protein constituent of lung surfactant - plays a dual role. It aggregates DPPC lipid, a major component of lung membrane, into a lattice-like structure that prevents the lungs from collapsing. SP-A is also known to recognize and bind bacterial lipids, namely lipid A, on surfaces of gram-negative bacteria, thereby helping to initiate various clearance mechanisms. However, it was unclear how SP-A exhibits such functional duality with its binding to two different types of lipids. A recent study used molecular dynamics simulations with NAMD to unravel the dual role of SP-A. Combined with crystallographic and mutational analyses, researchers have discovered several critical, non-canonical lipid binding sites that involves cation-π interactions and hydrogen bonds. Simulations have also revealed that SP-A binds stronger to bacterial lipid (lipid A) than to surfactant lipid (DPPC lipid), which suggests SP-A may prioritize its host defense functions by transferring from lung membrane to bacterial surface. These findings in atomistic detail will enable experimentalists to enhance the antimicrobial function of SP-A. More on our lung surfactant protein website.
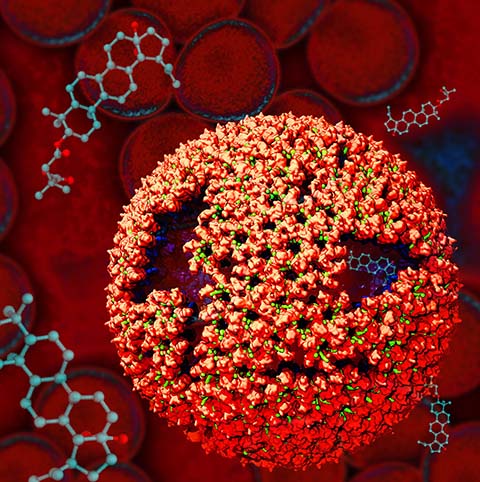
image size:
0 bytes
made with VMD
Virus capsids, specialized protein shells that encase the genome of viral pathogens, play critical roles in regulating viral infection, and are, thus, of great pharmacological interest as drug targets. In particular, small-molecule drugs (typically <900 Da) represent a promising antiviral strategy, and a number of such compounds have been developed to inhibit virus capsids by interfering with their assembly and uncoating processes. Importantly, to explain the mechanisms by which drugs disrupt capsid function, as well as to successfully design novel therapeutics, the interactions of drug molecules with their capsid targets must be studied at full chemical detail. Toward this end, all-atom molecular dynamics simulations are emerging as an essential technique to investigate the effects of small-molecule drugs on capsid structure and dynamics. Research presented in a recent Perspective applying simulations in NAMD to study drug-bound hepatitis B virus (HBV) and human immunodeficiency virus type 1 (HIV-1) capsids suggests the types of valuable chemical and physical information computational approaches can reveal, and underscores the importance of simulating, not isolated capsid proteins, but functional assemblies up to the level of complete capsids. Notably, through analysis with VMD, the study found that binding of the drug HAP1 to the HBV capsid causes global structural changes that subtly alter the overall capsid shape, including a flattening of capsid curvature. Further, the study found that the binding of the drug PF74 to the HIV-1 capsid imposes rigidity and causes shifts in allosteric communication pathways connecting distant regions of the capsid protein. The authors of the Perspective anticipate that many other such exciting discoveries regarding virus capsid function and their use as drug targets lie just ahead on the horizon, and molecular dynamics simulations will drive these discoveries pending a series of notable advancements in computational methodology.
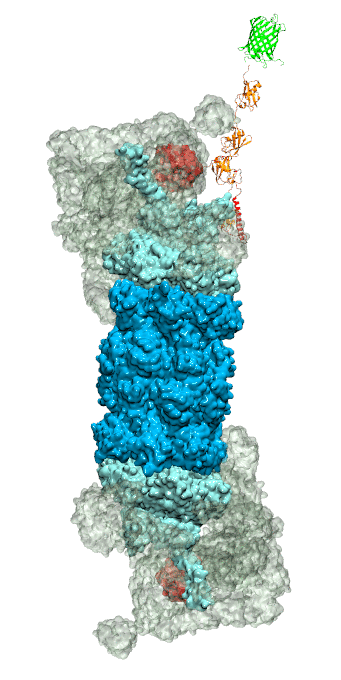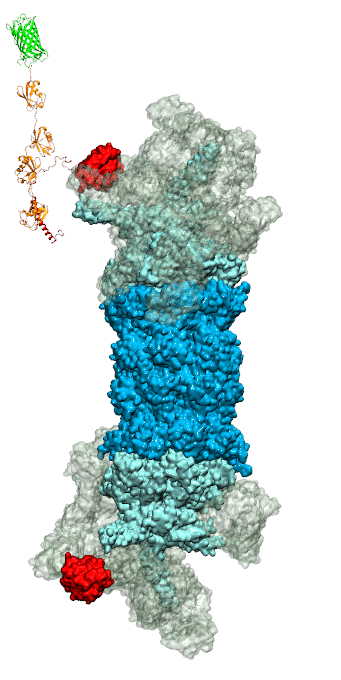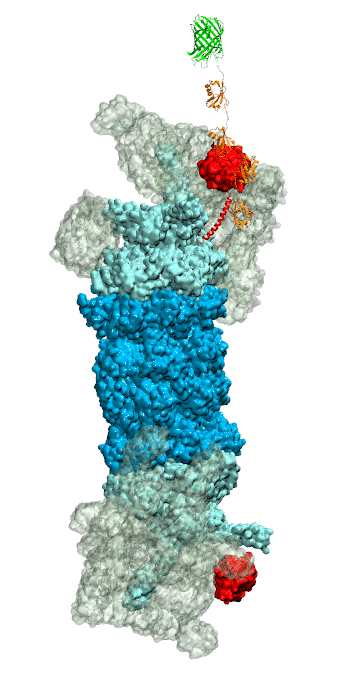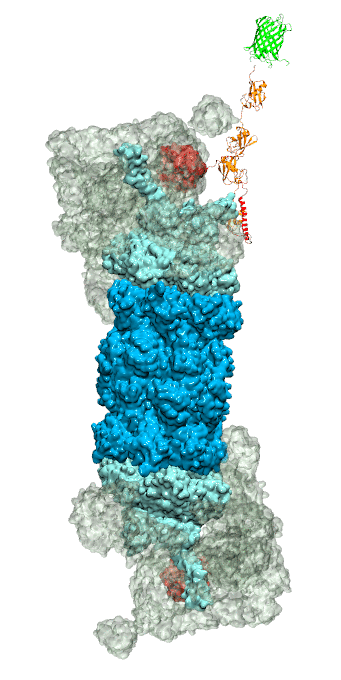
image size:
711.1KB
made with VMD
While waste recycling became popular in our daily life, living cells mastered waste recycling of their protein content since their very beginning. Recycling of unneeded protein molecules in cells is performed by a molecular machine called 26S proteasome, which cuts these proteins into smaller pieces and releases the pieces into the cell interior for reuse as building blocks for new protein. Proteins that need to be recycled are usually those that are misfolded. Proteins are recognized as such by the cells' so-called quality control system. This system labels misfolded proteins by a tag made of tetra-ubiquitin protein chains. The 26S proteasome machine recognizes and binds to these tags via its subunit Rpn10. After Rpn10 binds to the tetra-ubiquitin tag and pulls the protein close, the 26S proteasome unwinds the tagged protein and cuts it into pieces. A recent study, based on molecular dynamics simulations with NAMD, sheds light onto how 26S proteasome and Rpn10 recognize the tetra-ubiquitin tag in three stages: In stage 1 of the recognition process conserved complementary electrostatic patterns of Rpn10 and ubiquitins guide protein association; stage 2 induces refolding of Rpn10 and tetra-ubiquitin tag; stage 3 facilitates formation of hydrophobic contacts between the tag and Rpn10. More information is available on our 26S proteasome website.
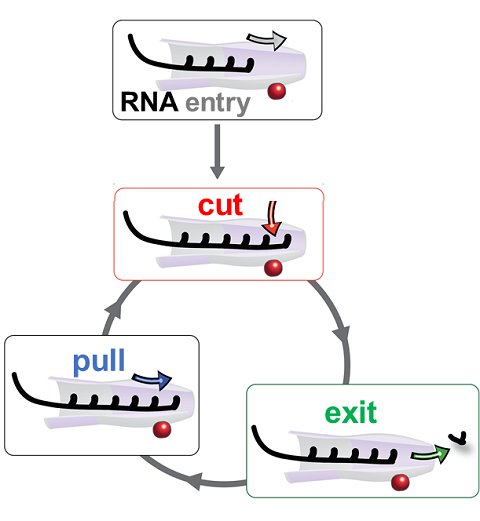
image size:
386.7KB
made with VMD
RNA molecules are continuously synthesized in living cells as carriers of biological information written in the sequence of basic RNA units, called nucleotides. To keep cells healthy, RNA molecules not longer needed or with errors have to be removed. A large barrel-like protein complex, the RNA exosome, is a molecular machine that degrades unneeded RNA molecules, pulling them inside its long internal channel and cutting them sequentially into single nucleotides. A new molecular dynamics study, employing NAMD, shows that a special active protein subunit of the exosome, called Rrp44, grips tightly the RNA molecule throughout its extended channel. Rrp44 grips RNA molecules with five or more nucleotides in length while their ending nucleotides get sequentially cut, whereas shorter RNAs are only weakly bound and unlikely to be cut. The simulations reveal how the exosome can act both as a molecular motor that pulls RNA, without energy input other than the one released in nucleotide cleavage, as well as an enzyme that cuts RNA. More information is available on our RNA exosome website.

image size:
101.5KB
made with VMD
When experimental-computational biologists embarked on the great challenge of resolving the atomic level structure of the HIV virus capsid that contains the virus' deadly genetic cargo, they were advised by referees not to try as the capsid is too big, too irregular, and nobody would need the highly resolved structural information. Stubbornly, the researchers went ahead anyway and succeeded getting the atomic resolution structure, overcoming size and irregularity challenges (see highlight Elusive HIV-1 Capsid). But the question remained: Is the atomic level structure of the huge HIV capsid made of 1,300 proteins useless? The HIV capsid is a closed container made of protein pentamers and hexamers, with a surface of continuously changing curvature. Two recent experimental-computational studies demonstrate now that the capsid structure is far from useless, in fact, it is a great treasure. The first study was published last year and showed that the human protein, Cyclophilin-A (CypA), involved in several diseases, interacts with the HIV capsid and affects the capsid's dynamic properties (see highlight HIV, Cells and Deception). In a second, recent study, guided by cryo-EM measurements and benefiting from large-scale molecular dynamics simulations with NAMD, researchers could resolve with new accuracy the binding of hundreds of CypA proteins on the capsid's surface. They found that CypA binds along high curvature lines of the capsid, which enhances stiffness and stability of the capsid, even though only about half of the capsid is actually covered by CypA. The limited levels of CypA stabilize and protect the viral capsid as it moves through the infected cell towards the cell's nuclear pore where nuclear proteins additionally bind to the capsid at places not covered by CypA and promote there uncoating and release of the capsid cargo into the nucleus. More information is available on our retrovirus website and in a news release.
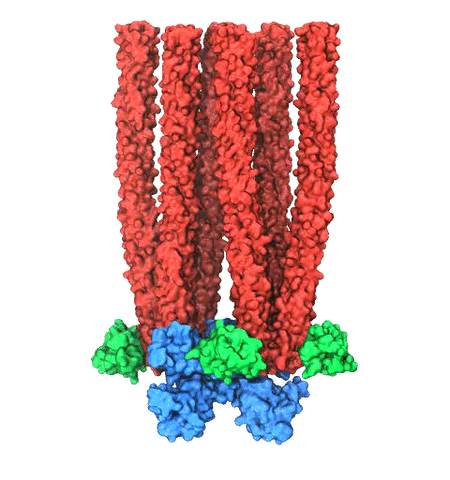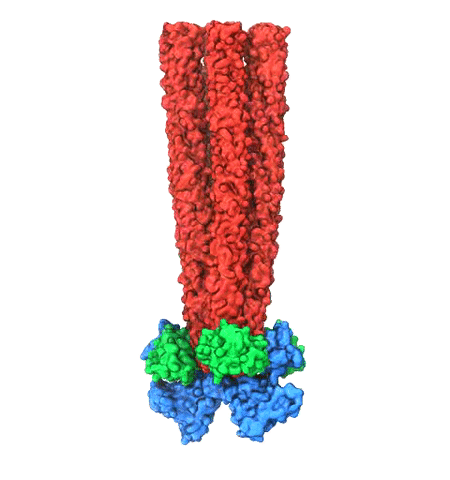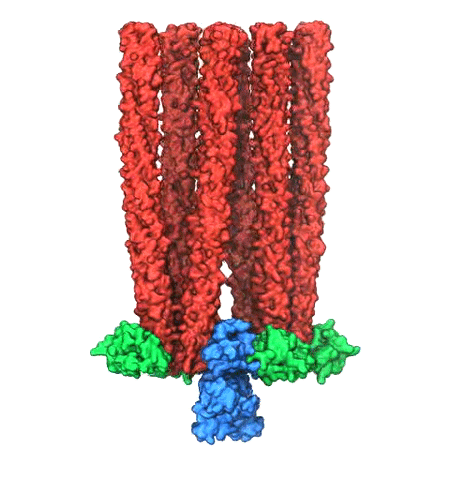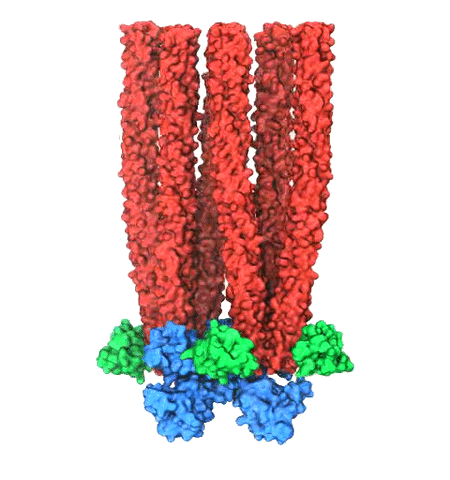
image size:
642.1KB
made with VMD
Motile bacteria position themselves within their habitats optimally, seeking proximity to favorable growth conditions while avoiding unfavorable ones. Cues used for this placement come in the form of small chemicals, so-called attractors and repellants, as well as physical factors such as favorable visible light and unfavorable UV radiation. To balance such a broad range of factors, bacteria monitor their environments and respond by way of a fundamental sensory capability known as chemotaxis. Chemotactic responses in bacteria involve large complexes of sensory proteins, known as chemosensory arrays, that process the information obtained from the bacteria's habitat to determine its swimming pattern. In this sense, the chemosensory array functions as a bacterial brain, transforming sensory input into motile output. Despite great strides in the understanding of how the chemosensory array's constituent proteins fit and work together, a high-resolution description of the kind needed to explore in detail the molecular mechanisms underlying sensory signal transduction within the array has remained elusive. A new study, utilizing cryo-electron microscopy and molecular dynamics simulations with NAMD, reports the highest resolution images yet of the bacterial brain's molecular anatomy. Using computational techniques, structural data from X-ray crystallography and electron microscopy are compared to derive an atomically resolved model of the chemosensory array's extended molecular structure that involves millions of atoms. Subsequent simulations of the model revealed a novel conformational change in a key sensory protein, that is interpreted as a key signaling event in the translation of chemosensory information into swimming pattern. More details on this work can be found in a recent news release as well as on our bacterial chemotaxis website.
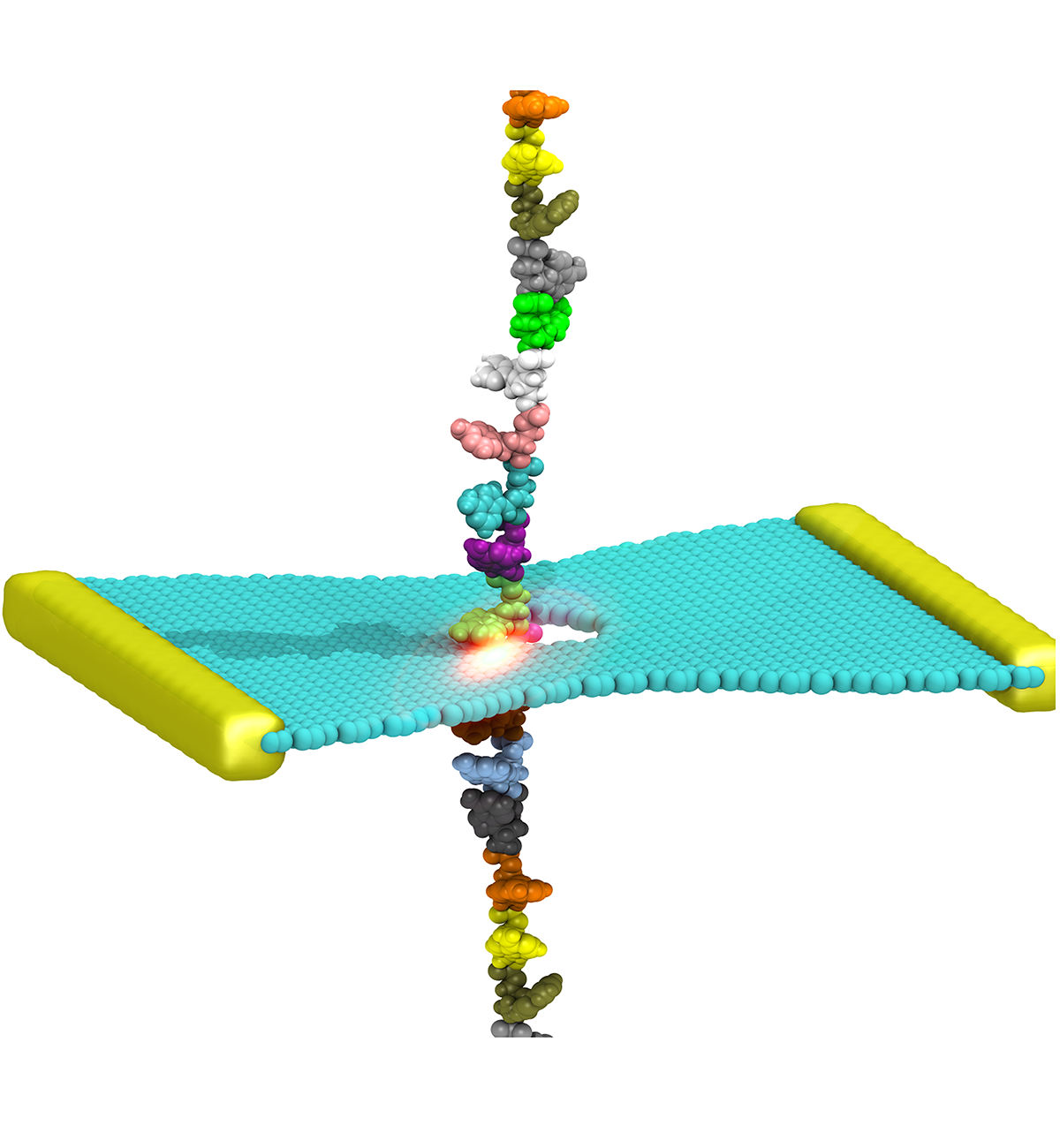

image size:
1.7MB
made with VMD
DNA sequencing is achieved by following a strand of DNA at a speed that permits recognition of the DNA bases in their actual order, thousands of bases or more for each pass. Nanotechnology can assist in the task, in principle, by furnishing sensors that can resolve single DNA bases and nano-mechanical actuators that pull the DNA in a controlled fashion passing through the sensor. Instead of building and testing actual sensors and actuators it is cheaper and faster to simulate them. Nanoengineers have indeed succeeded with such simulations over the last decade focussing on silicon technology (see October 2004 highlight: Transistor Meets DNA). Now the engineers have moved with their simulations to graphene technology that promises much better resolving power as sensors are thinner and as signals can be detected electronically in graphene (December 2013 and November 2014 highlights). The main unsolved problem is the mechanical actuator: how can one control movement of DNA through a graphene sensor such that measured signals become less noisy and bases can be recognized? A recent study, based on molecular dynamics simulations with NAMD and quantum electronics calculations of graphene, suggests use of an actuator that simultaneously stretches the DNA and pulls it through the sensor. This manipulation leads to a stepwise translocation of DNA through the graphene nanosensor, slowing down DNA translocation and stabilizing DNA bases inside the sensor. Read more on our graphene nanopore website.
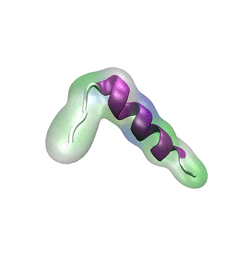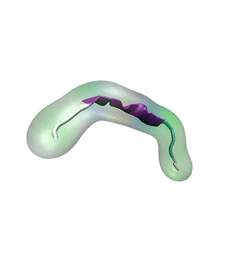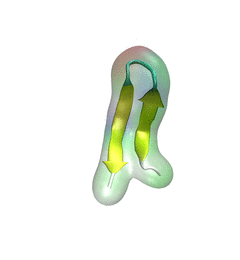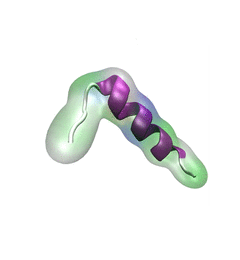
image size:
768.0KB
made with VMD
For centuries, millions of people around the globe have been troubled with a movement disorder that usually starts with a tremor in one hand. The disorder, later known as Parkinson's disease, affects commonly older individuals and disrupts patient's movement, sleep and speech from the brain. There is currently no cure for the disease. Key to the disease, progressively occurring in patient's brain, is the loss of neuron cells due to aggregation of a small protein named α-synuclein. Extensive studies have been carried out, yet the function of the protein remains a mystery. It is amazing that aggregation of such small proteins eventually leads to neuronal cell death and generates tremendous difficulties in peoples' life. In a recent report, a team of computational scientists attributed the cause of α-synuclein aggregation to a hairpin structure involving just a small region (amino acids 38-53) in the middle of the protein. With extensive simulations (over 180 μs in total), the researchers revealed that a short fragment encompassing region 38-53, exhibiting a high probability of forming a β-hairpin structure, is a key region during α-synuclein aggregation. Moreover, the researchers predicted a mutation that impedes β-hairpin formation, thereby retarding α-synuclein aggregation. The discoveries, made possible through the software NAMD and VMD, are expected to shed light on the mechanism underlying Parkinson's disease and to inspire the design of drugs. More on our α-synuclein website.
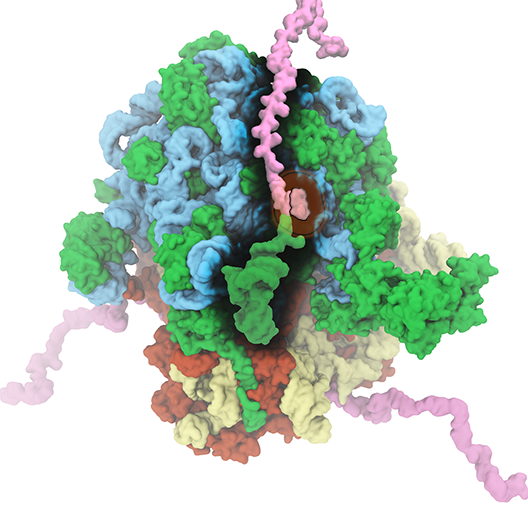
image size:
816.0KB
made with VMD
The ribosome is the ubiquitous machine in all living cells responsible for translating the cell's genes into functional proteins. The majority of antibiotic drugs target the ribosomes of bacterial cells while leaving human ribosomes unharmed. An example are the most widely-prescribed antibiotics, erythromycin and telithromycin. They kill bacteria by changing the properties of bacterial ribosomes and, thereby, disturb the bacterial protein production (see the Oct 2014 highlight Antibiotic Action on the Ribosome). However, modern bacteria fight antibiotic drugs; exposing them to a specific kind of antibiotic drug for too long will trigger the expression of drug-resistance genes, which protect the bacteria, eventually making the drug useless. Due to historical overuse of antibiotic drugs, clinic antibiotic drugs have experienced today serious drug-resistance problems. In a joint effort of computational and biomedical investigations, reported recently, molecular dynamics simulations with NAMD and systematic mutation experiments showed that the above antibiotics interact in a bacterial ribosome with a drug resistance gene - coded nascent protein and make it stall translation; however, engineered simple mutations in the bacterial gene can abolish stalling and, thereby, prevent the effect of drug resistance genes. The research suggests that engineered mutations might be a strategy to prevent antibiotic resistance. Read more on our Ribosome website.



