Highlights of our Work
2026 | 2025 | 2024 | 2023 | 2022 | 2021 | 2020 | 2019 | 2018 | 2017 | 2016 | 2015 | 2014 | 2013 | 2012 | 2011 | 2010 | 2009 | 2008 | 2007 | 2006 | 2005 | 2004 | 2003 | 2002 | 2001
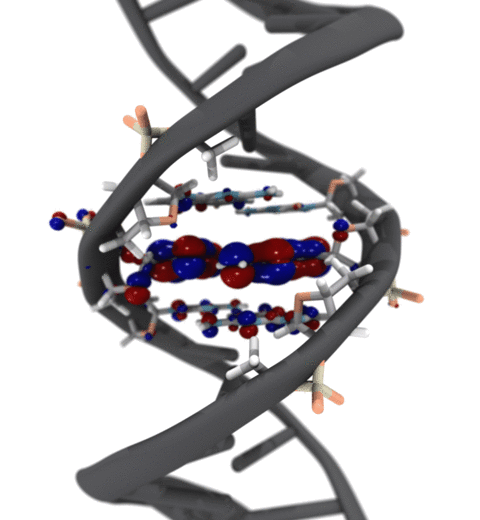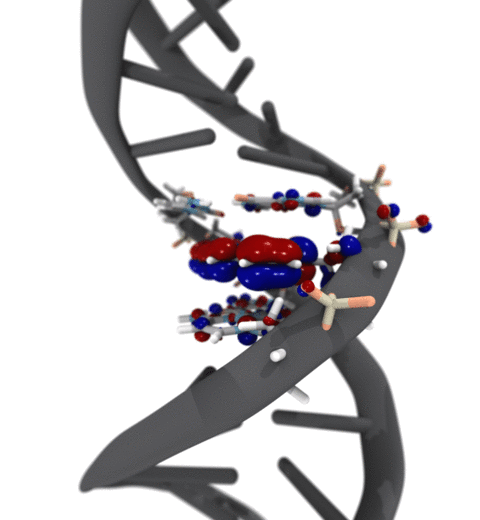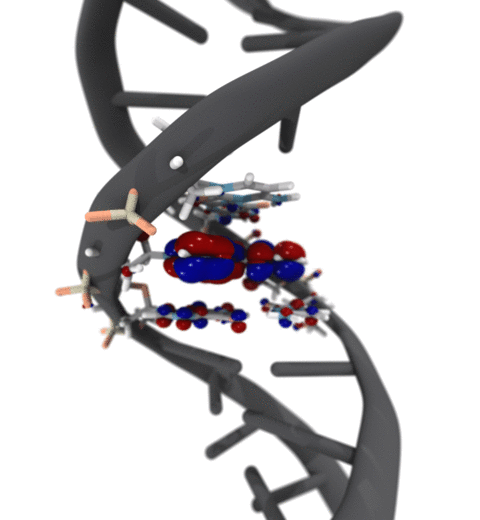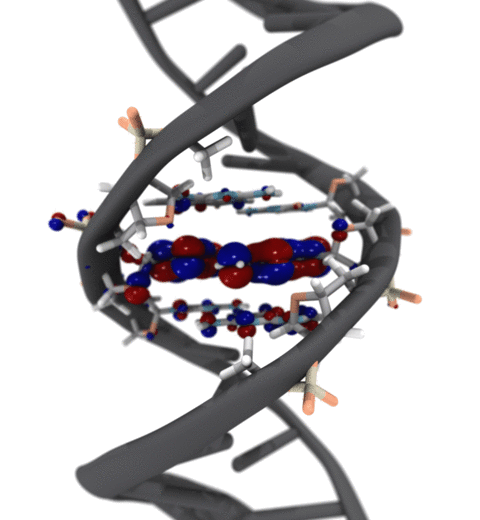
Quantum mechanics (QM) is crucial to investigate subatomic mechanisms that occur in biology. However, studying entire biomolecular systems quantum mechanically is computationally prohibitive. Traditionally, NAMD employs classical molecular mechanics (MM) to determine the movement of a molecule, solving Newton's equations of motion and treating atoms as spheres and bonds as springs. Combining both approaches, hybrid QM/MM methods employ the quantum mechanics formalism to key regions of the biological system, while using molecular mechanics approach to include the effects of the surrounding area. NAMD's new QM/MM interface can be combined with many molecular dynamics protocols, such as enhanced sampling and free energy calculations. This combination was crucial in the investigation, appearing in Nature Methods, of the mechanism that sets the genetic code, when DNA's three-letter code is translated to protein's amino acid residues. Uniting a large set of tools in VMD, QwikMD, and NAMD, the QM/MM simulations revealed the subatomic details of this mechanism, providing an unique view for this essential step of life. Read more in Nature Methods, or from our QM/MM website.
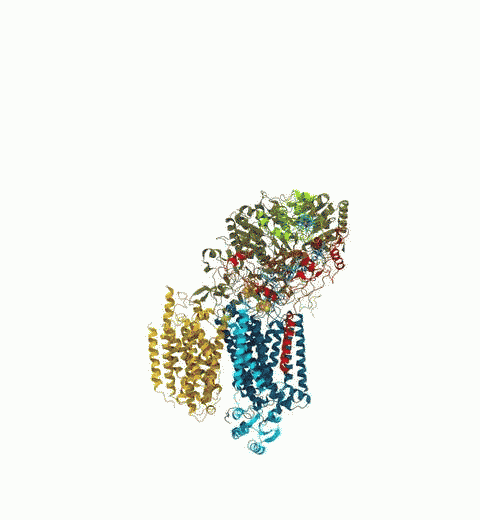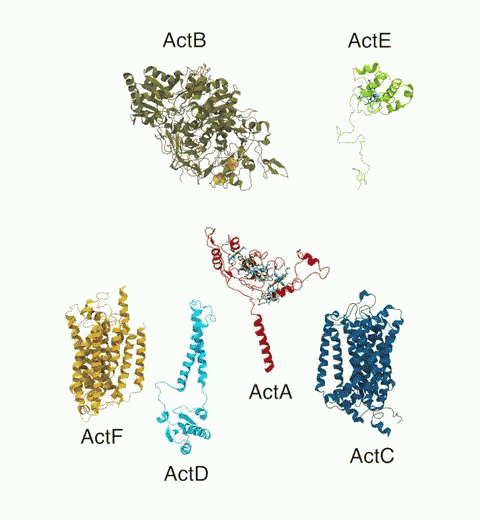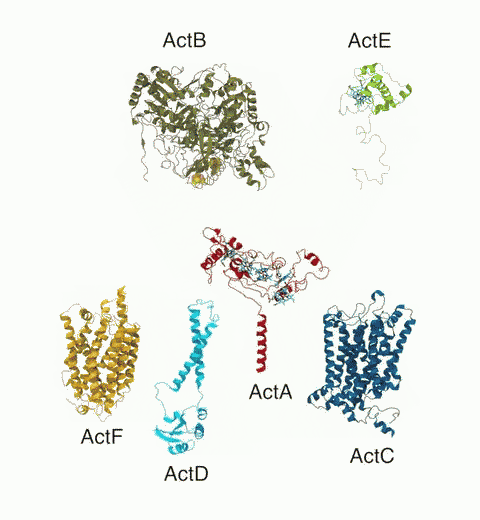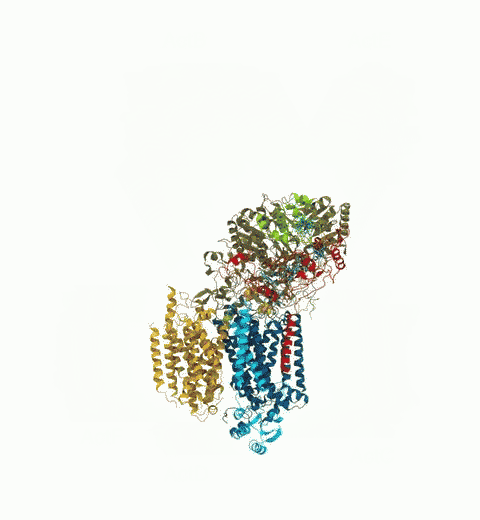
image size:
1.9MB
made with VMD
Bacteria are remarkable at tailoring their bioenergetic machineries to adapt to and thrive in diverse and ever-changing environments. A critical metabolic task is the efficient extraction of energy from food. Similar to us, many bacteria pass electrons to oxygen or other acceptors with the help of membrane-bound enzymes, and in doing so, they move protons across their membrane, thus generating a transmembrane voltage (much like a battery) that can be used for ATP synthesis. Typically, one enzyme passes an electron to its downstream enzyme via random collisions. However, sometimes these enzymes can form a supercomplex that positions the enzymes very close to each other and with the right pose for faster electron transfer and therefore more efficient energy conversion. In a recent paper reporting a three-way collaboration between biochemists, experimental structural biologists, and the Center researchers, the structure of one such supercomplex (termed Alternative complex III) was determined at an atomic resolution through a combination of cryoEM and advanced molecular modeling and simulation tools performed with NAMD and VMD. Read more about this study in Nature.
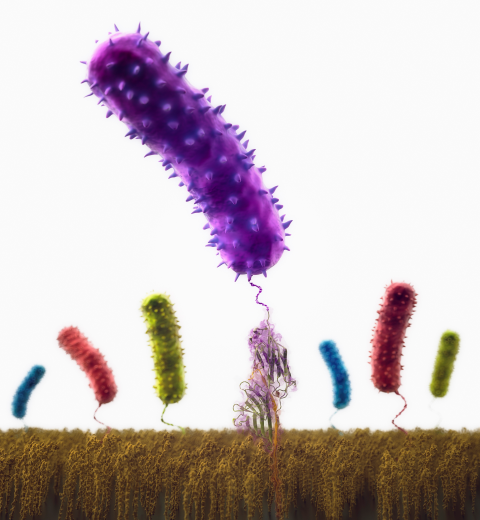
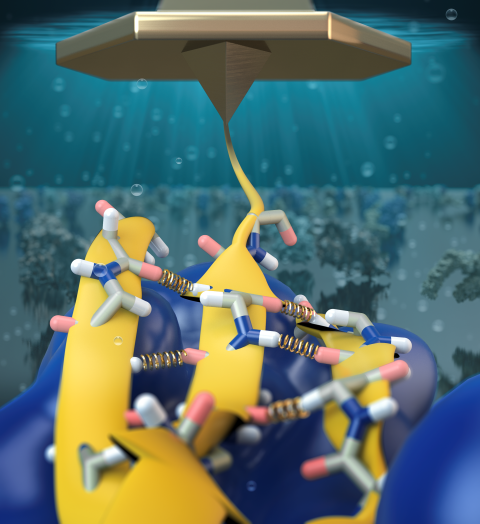
Staph infections are caused by bacteria commonly found on the skin of healthy individuals, where they can only cause relatively minor skin problems. However, when staph bacteria enter a person's bloodstream, they can travel to locations deep within the body causing infections that are often hard to treat. Not surprisingly, staph infections are the leading cause of healthcare-related, so called nosocomial infections. Particularly vulnerable areas are medical devices such as artificial joints or cardiac pacemakers, where the bacteria strongly stick to through formation of biofilms. Central to biofilm formation is an unusually tight interaction between microbial surface proteins called adhesins and the extracellular components of the host cells. In a recent report in Science , researchers used a combination of atomic force microscopy and GPU-accelerated molecular dynamics simulations using QwikMD and NAMD , to explore how the connection between adhesin and its target fibrinogen peptide can withstand huge forces greater than 2 nano-Newtons (see also Perspective in Science. ). The geometry and molecular details of the interaction ensures that, when pulled, the load is distributed over many hydrogen bonds all of which need to be broken at once before separating the bacterium from the surface of the host cell (see video on YouTube ). The unexpected, new mechanism, expands our understanding of why pathogen adhesion is so resilient and open new avenues for an intelligent design of antimicrobial therapies through development of anti-adhesion drugs. Read more in Science.
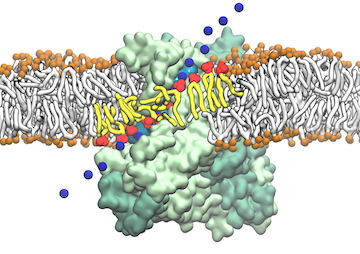
image size:
128.0KB
made with VMD
Proteins play major mechanical roles in a cell. Mechanical properties of proteins can be substantially modulated by slight changes in their amino acid composition, an ability essential to bacterial cells, which use protein repeats with small modifications to tune their mechanical strength, and thus their functional properties. A prime example of the role of biomechanics in the cell is found in symbiont gut bacteria who need to hold to their food source in such turbulent environments as the rumen of the cow. Proteins evolved for this purpose are known actually to be among the strongest mechanical biomolecules. Key to the process are molecular tentacles, so-called cellulosomes, on the surface of symbiotic bacteria. The cellulosomes develop a tight grasp on and then effective cleavage of hardy cellulose fibers of the grass. In a joint experimental-computational study researchers investigated the mechanical properties of cohesin, a major cellulosome component, under force. Using NAMD, all-atom steered molecular dynamics (SMD) simulations on homology models offered insight into the process of cohesin unfolding under force. Based on the differences among the individual force propagation pathways and their associated correlation communities, the researchers were capable of designing protein mutants to tune the mechanical stability of the weakest cohesin. The proposed mutants were tested with high-throughput atomic force microscopy experiment revealing that in one case a single alanine to glycine point mutation suffices to more than double the mechanical stability. Read more about our cellulosome research here.