Highlights of our Work
2026 | 2025 | 2024 | 2023 | 2022 | 2021 | 2020 | 2019 | 2018 | 2017 | 2016 | 2015 | 2014 | 2013 | 2012 | 2011 | 2010 | 2009 | 2008 | 2007 | 2006 | 2005 | 2004 | 2003 | 2002 | 2001

image size:
621.3KB
made with VMD
Sunlight powers life on Earth. This basic fact has been known since ancient times and retold many times in many cultures. Today, scientists understand the means through which light powers life at atomistic detail, as a chain of processes climbing scales from electronic interactions to cooperation between proteins to cell-scale integration of energy conversion. These processes have been illustrated in a recent video
that was awarded the
BEST SCIENTIFIC VISUALIZATION OF 2019
at the SC19 conference, Scientific Visualization & Data Analytics Showcase.
The video is a joint production with the team of
Donna Cox
at
NCSA
and is based on a decade long collaborative effort with the experimental group of
Neil Hunter
and a lifetime research interest for
Klaus Schulten.
The underlying scientific investigation illustrated in the video
was presented in
18 manuscripts over the past decade,
from
atomic scale structural modeling
to
organelle-scale
and
cell-scale
integration of function.
These modeling efforts also led to
a molecular dynamics simulation of the chromatophore
using NAMD,
recently published in
Cell.
The video segment,
produced with VMD,
that won as the Best Scientific Visualization is an excerpt from
the fulldome movie
'Birth of Planet Earth' released to planetariums worldwide by Spitz Creative Media.
The oldest story of humanity — light powering life — coming soon to a theater near you.
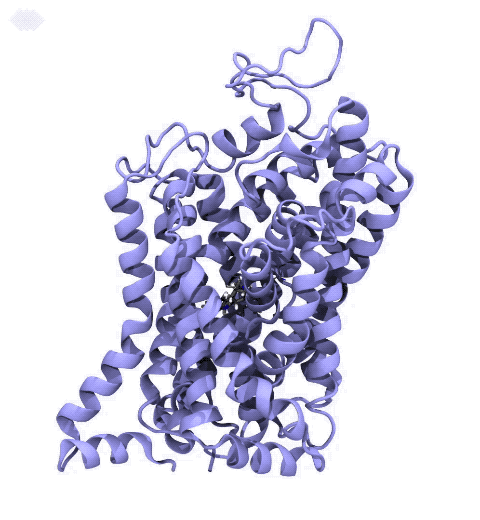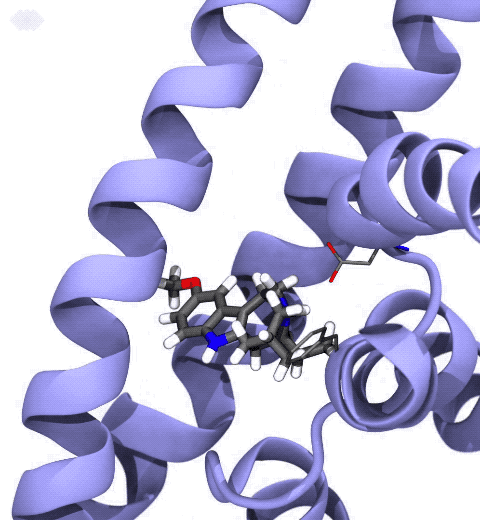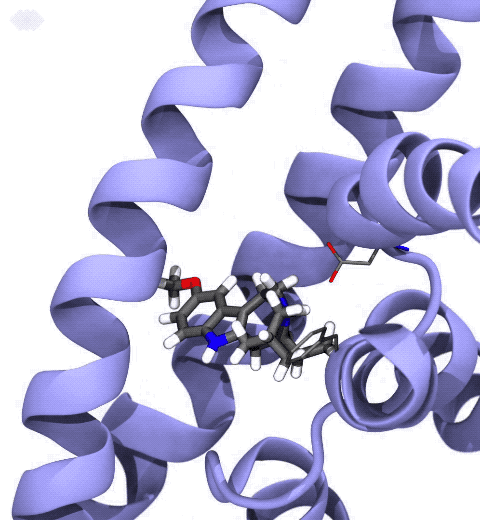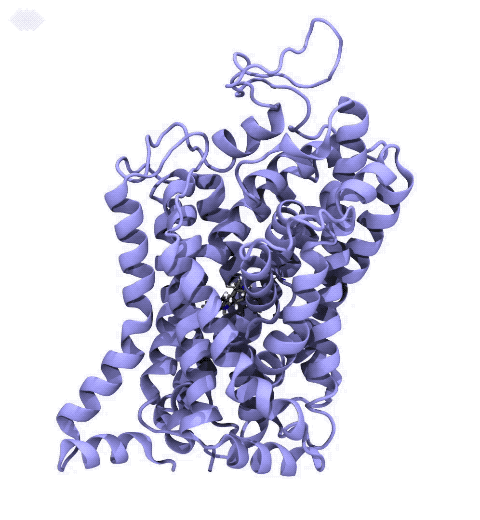
image size:
7.7MB
made with VMD
Neurotransmitters are chemical messengers diffusing across the synapse to propagate nerve impulses. The serotonin transporter (SERT) is a protein embedded in the cellular membrane of the pre-synaptic neuron, which recycles the neurotransmitter serotonin. It plays pivotal roles in modulating
behavior, rendering it a primary drug target for neuropsychiatric disease. Inhibition of the serotonin uptake by SERT contributes to the mechanism of clinically used antidepressants. To provide structural insights into the transport mechanism, our collaborators in the Gouaux lab (OHSU) captured a series of structures of SERT in multiple functional states using cryoEM. The structures were co-crystalized with an inhibitor, ibogaine, a hallucinogenic drug found in plants with anti-addictive potential, in the substrate-binding pocket. However, the exact binding poses of ibogaine in these structures remained ambiguous due to the limited resolution, prohibiting a direct interpretation of ibogaine's inhibition mechanism. To settle this problem, we developed a computational docking approach to systematically search for the most probable ibogaine binding poses, which best fit into the experimental density maps and exhibit considerable stability in the following by molecular dynamics simulations performed with NAMD. Read more in Nature.
Modern molecular simulation, visualization, and modeling techniques often require high-performance computing hardware
to obtain the best efficiency. However, access to such hardware can be a barrier for some researchers.
As an alternative, cloud computing provides a cost-effective and practical solution for many molecular modeling tasks and for small and moderate size molecular dynamics simulations. The cloud computing model
provides researchers with access to powerful computational equipment
that would otherwise be too costly to procure, maintain,
and administer on their own. Additionally, cloud platforms can easily bundle different software packages
used in a modeling workflow to guaruntee their availablity and
interoperability on a standardized system.
We have adapted our molecular modeling applications NAMD, VMD, and associated tools to
operate within the Amazon Web Services (AWS) Elastic Compute Cloud (EC2) platform, enabling popular
research workflows to be run remotely by scientists all over the globe,
with no need for investment in local computing resources and
a reduced requirement for expertise in high performance computing technologies.
Recent advancements in GPU virtualization technology now make it possible to use cloud computing for
large-scale scientific computing, data analysis, and visualization tasks. The latest release of our
cloud software can be run on EC2's latest graphics hardware,
backed by DCV, to provide a smooth and seamless
graphical working environment. Instance types specially designed for parallel CPU or GPU workloads
provide users with access to the hardware they need to run even the most demanding features of NAMD and VMD,
all from their much less powerful personal computers and even laptops. These instances can be paid for on an on-demand, as-needed basis, and some
workflows, such as Molecular Dynamics Flexible Fitting, can be run for less than the cost of a cup of coffee.
Our cloud virtual machine image is availble in the Amazon Marketplace,
which provides users with
a very simple one-click launch using pre-configured choices
of instance types that have been tested with our software.
To learn more,
see our
cloud research page.




