VMD 1.9 contains many new and improved features for generating
high quality renderings of molecular graphics and for creation
of movies of both static structures and molecular dynamics simulation
trajectories.
The molecular graphics export features of VMD have been
significantly enhanced in the latest version.
The
built-in Tachyon parallel ray tracer in VMD has
been improved with faster rendering, and it now supports all
of the special VMD rendering and shading features including
angle-modulated tranparency, all depth cueing modes,
and background gradients.
VMD now supports the X3D scene format, which can be used to make
documents with interactive embedded
3-D molecular graphics. The latest WebGL-capable web browsers can
interactively display VMD molecular graphics
in X3D format, without the need for any additional browser plugins.
The VMD export modules for Raster3D, RenderMan, POV-Ray, Tachyon,
VRML2/VRML97, and X3D have been udpated to add full support for
text rendering, e.g. atom labels, angles, bond lengths,
time-varying distance measurement labels, and user-defined text,
enabling text labels to be shown in high-quality ray traced
renderings and movies.
VMD now exports large models to the Wavefront OBJ scene format
with much greater efficiency. VMD now exports material properties,
molecular representation groupings, and exports molecular geometry
in a more efficient way, resulting in much smaller scene files that are
easier for other programs to work with.
The Wavefront OBJ file format supported by
many commercial rendering and animation packages, and with these improvements
professional visualization artists can more easily incorporate molecular
graphics directly from VMD into their rendering tools. The improved
Wavefront OBJ export feature has been extensively tested with the most
recent versions of
Autodesk Maya,
and the embedded VMD representation "grouping" information
makes it easier to work with imported VMD scenes within the
Maya graphical interfaces.
The
VMD movie maker plugin
has been updated to support a broader range of rendering and movie
compression formats and options.
The new
ViewChangeRender
plugin provides an easy-to-use graphical interface for managing multiple
VMD camera viewpoints and making movies that fly the camera between
multiple viewpoints.
The new version of VMD adds support for a broader range of stereoscopic
display hardware, including displays that use "checkerboard" and
column-interleaved formats used by DLP HDTVs and projectors, and
the column-interleaved format used by autostereoscopic displays.
The anaglyph (red/cyan) stereo mode has been adapted to support
inexpensive gaming GPUs and laptop graphics chipsets.
New graphical menus make it easy to swap the left and right eyes
in all supported stereo modes, for hardware configurations that
reverse the eyes, and to simplify the creation of
cross-eye and wall-eye stereo images.
VMD 1.9 further advances the use of
GPU accelerated molecular visualization and analysis,
based on
NVIDIA CUDA,
and most recently adding support for
OpenCL
(use of OpenCL requires compilation from source code at present).
As
reported in several publications,
the massively parallel architecture of GPUs makes them ideal devices
to accelerate many of the computationally demanding calculations
in VMD. The range of acceleration provided by GPUs depends on the
capabilities of the specific GPU device(s) installed, and the details
of the calculation. Typical acceleration factors for the algorithms
in VMD on a single high-end GPU are:
electrostatics 22x to 44x,
implicit ligand sampling 20x to 30x,
calculation of radial distribution functions 30x to 90x,
molecular orbital calculation 100x to 120x.
Details on making best use of the GPU acceleration capabilities
in VMD are
provided here.
VMD 1.9 incorporates many improvements aimed at increasing
its structure building capabilities.
The latest version of the
molefacture plugin
performs geometry optimisations and can assign charges using
the latest versions of SQM and antechamer (distributed with ambertools).
Molefacture now allows structures to be built for the OPLS force field,
based on included CHARMM-formatted OPLS parameter files.
Molefacture also provides an FEP setup tool to aid in the generation of
structures and input files for free energy perturbations with NAMD.
VMD reads, stores, and writes angles, dihedrals, impropers, and
cross-term maps, and includes text commands for querying and setting
these fields enabling the development of flexible structure
building tools such as
the
new topotools plugin.
The toptools plugin makes it much easier to develop
customized structure building tools like the newly updated
Carbon Nanostructure Builder plugin.
The new
Chirality, and
Cispeptide plugins help
researchers build their structures, by detecting, visualizing, and
correcting, and enforcing chirality and peptide bond configuration
in proteins. A new tutorial describing the Cispeptide and Chirality
plugins is
available here.
The new
MDFF plugin provides
commands for fitting atomic structures to a density map, using the
molecular dynamics flexible fitting method.
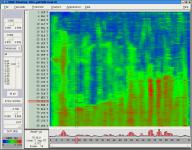
Trajectory analysis with Timeline
VMD 1.9 also includes several new molecular dynamics analysis features.
The new
ParseFEP plugin
provides a set of tools for analyzing free-energy perturbation (FEP)
calculations performed with NAMD.
The
PLUMED plugin
allows collective variable analysis from within VMD, using PLUMED.
VMD includes a new GPU-accelerated method for computing radial distribution
functions up to 90x faster than with the CPU alone, and a new version
of the graphical
g(r) plugin
includes support for the GPU-accelerated calculations.
The updated
Timeline plugin
provides an interface for viewing temporally changing per-residue
attributes of a molecular structure. It can also display temporally changing
attributes of a set of VMD selections, for example a set of all the
salt-bridge pairs observed in a trajectory.
The controls allow selection of the molecule, or part of the molecule,
used for the calculation.
The graphical display of residues and timesteps can be scrolled and
zoomed as necessary to see results for large structures and long trajectories.
The latest version improves performance for large structures and long
trajectories, provides more analysis functions and options,
and significantly improves the printing features for saving Timeline
trajectory analysis plots as encapsulated Postscript (.eps) files.
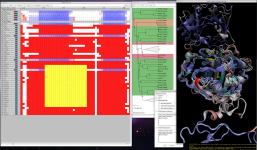
MultiSeq analyzes large datasets efficiently
This release of VMD includes the newly revised
MultiSeq plugin version 3.1.
MultiSeq has added new support for
MAFFT for multiple sequence alignments.
RMSD and Q calculations now work for RNA as well as DNA.
The title shown for a sequence can now be changed from simply 'Sequence Name'
to a variety of options including Scientific Name, Common Name,
Domain of Life, and many others.
Major efforts have been directed toward improving the ability of MultiSeq
to handle large data sets, and the new MultiSeq is capable of loading
and analyzing 100,000 sequences on a typical desktop machine.
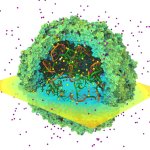

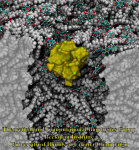

 VMD/Tachyon renderings with
VMD/Tachyon renderings with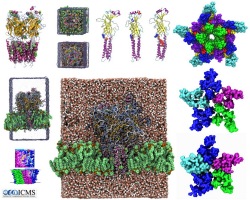 VMD molecular scenes with ambient
VMD molecular scenes with ambient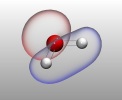
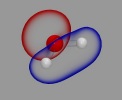 Angle-modulated transparency in VMD:
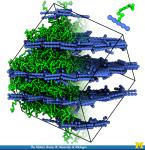
Angle-modulated transparency in VMD: VMD 1.9 exports large models to Wavefront OBJ
VMD 1.9 exports large models to Wavefront OBJ