MultiSeqMultiSeq is a unified bioinformatics analysis environment that allows one to organize, display, and analyze both sequence and structure data for proteins and nucleic acids. Special emphasis is placed on analyzing the data within the framework of evolutionary biology. In any publication of scientific results based completely or in part on the use of MultiSeq, please reference:
Elijah Roberts, John Eargle, Dan Wright, and Zaida Luthey-Schulten. User's GuideVisit the Luthey-Schulten group MultiSeq home page for additional information and tutorials. |
 |
Download (Note: MultiSeq is located in the Extensions menu as Analysis->MultiSeq) |
Learn A tutorial entitled "Evolution of Biomolecular Structure" is available for learning how to use MultiSeq. |
|
|
|
||
Features |
||
 |
 |
|
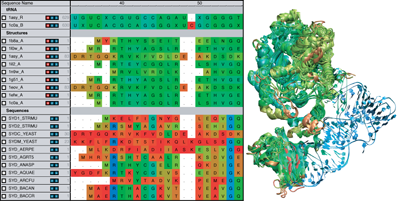
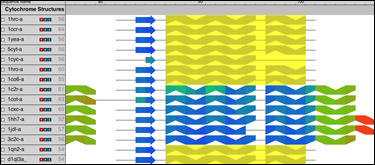
Combine Sequence and Structure data to gain insight into evolutionary changes in sequence, structure, and function. Compare sequence measures alongside structural measures with just a few mouse clicks. |
Quickly and Easily Import Data from many popular files formats or with an integrated BLAST search. Filter search results by sequence identity to eliminate duplicates or by taxonomy to focus a search. |
|
 |
 |
|
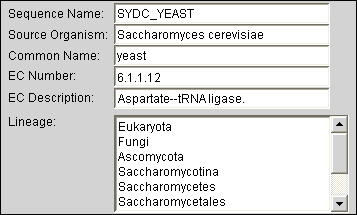
Automatically Find Metadata for sequences and structures from published databases. Use this information (such as taxonomic classification or enzymatic function) during all steps of the analysis process. View and edit metadata via the electronic notebook. |
Align Sequences and Structures using popular alignment algorithms: MAFFT or ClustalW for sequences and STAMP for structures. Perform multiple alignments or profile alignments on entire molecules or just select regions. |
|
 |
 |
|
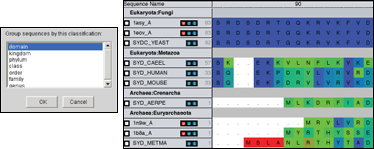
Organize Data into custom groupings to ease the process of analyzing large amounts of data. Automatically group by taxonomy to help put the data into an evolutionary context. |
Eliminate Bias and Redundancy by running the QR algorithms for sequence and structure. |
|
 |
 |
|
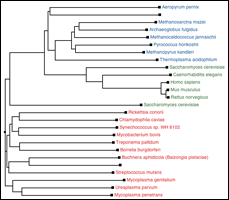
View and Manipulate Phylogenetic Trees representing your data by either computing distance trees on the fly or loading precomputed trees from other phylogenetic packages. |
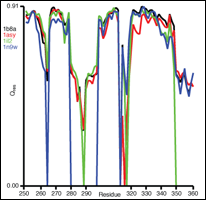
Plot Data Measures on a per residue basis with the built-in plotter. |
|
Plus... |
||
Work with Nucleic Acid sequences and structures using many of the same techniques as for proteins. |
||
Export Data to many popular file formats or create images of alignments, phylogenetic trees, or plots as publication quality PostScript graphics. |
||