VMD 1.8.7
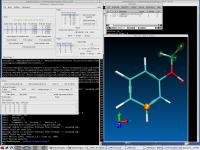
The Theoretical and Computational Biophysics Group is pleased to announce VMD version 1.8.7. VMD provides exciting new capabilities for the display of molecular orbitals arising in quantum chemistry simulations, graphical representations for illustration of carbohydrates and other multi-branched structures, high dynamic range images, and eerily "three dimensional" renderings using ambient occlusion lighting with production quality renderers. The MacOS X, Unix, and Windows versions of VMD now benefit from more extensive use of multiprocessor (and multicore) acceleration, greatly improving performance for several common structure analysis operations. By default VMD will now use all available processor cores to accelerate parallelized portions of the code which currently include several structure analysis routines, interactive molecular dynamics, and ray tracing. Many new and updated structure building and analysis tools have been added in this release, easing the process of setting up, running, and analyzing computer simulations of biomolecules. This release also contains many performance and efficiency improvements benefiting those loading, displaying, and analyzing large structures. In particular, this version of VMD makes extensive use of multi-core processors and GPU acceleration to speed up computationally demanding analysis and visualization tasks.
- VMD 1.8.7 Documentation, Release Notes, Tutorials
- Download VMD 1.8.7 for MacOS X, Unix, or Windows
- VMD 1.8.7 Development and Release History (large)
Major features included in VMD 1.8.7:
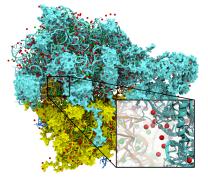
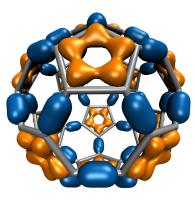
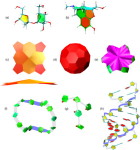
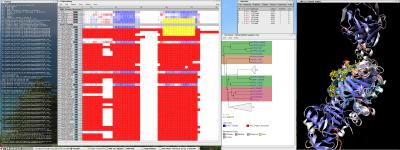
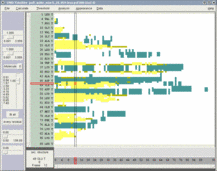
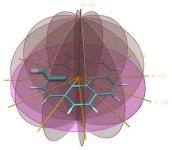
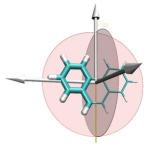
One of the key advancements included in VMD 1.8.7 is support for GPU accelerated visualization and analysis, based on NVIDIA CUDA. As reported in several publications, the massively parallel architecture of GPUs makes them ideal devices to accelerate many of the computationally demanding calculations in VMD. The range of acceleration provided by GPUs depends on the capabilities of the specific GPU device(s) installed, and the details of the calculation. Typical acceleration factors for the algorithms in VMD are: electrostatics 22x to 44x, implicit ligand sampling 20x to 30x, molecular orbital calculation 100x to 120x. Details on making best use of the GPU acceleration capabilities in VMD are provided here.Several new representations have been added to VMD in the new release. A new "Orbitals" representation allows the display (and animation) of molecular orbitals from quantum chemistry simulation log files. The new "Orbitals" representation benefits from GPU and multi-core CPU acceleration, allowing very high resolution orbital surfaces to be interactively rendered and animated, as reported here. The newly added "Polyhedra" representation can be used to render tetrahedra for various silica (SiO2) structures. The new "PaperChain" and "Twister" representations used to display molecules containing carbohydrate or other multi-branched structures, as reported here. The "PaperChain" representation highlights each ring in a molecular structure with a polygon that is colored according to ring pucker. The "Twister" representation traces glycosidic bonds with a ribbon that twists according to the relative orientation of successive sugar residues.This release of VMD includes the newly revised MultiSeq plugin version 3.0. Major efforts have been directed toward improving the ability of MultiSeq to handle large data sets, and the new MultiSeq is capable of loading and analyzing on the order of one hundred thousand sequences on a typical desktop machine. Additionally, MultiSeq can now correlate sequences and structures with other source of biological data, including the NCBI taxonomy databases, databases regarding microorganism growth temperatures, and enzyme function databases.VMD 1.8.7 incorporates many improvements aimed at increasing its structure building capabilities. VMD reads, stores, and writes angles, dihedrals, impropers, and cross-term maps, and adds new text commands for querying and setting these fields enabling the development of flexible structure building tools such as the new topotools plugin. The latest version of the molefacture plugin adds the ability for users to build and use custom fragments. The fragment libraries have also been expanded. The new version also includes an interface for antechamber, providing users with autotyping and semiempirical geometry optimizations for fast structure cleanup. Finally, a new graphical interface greatly simplifies the process of setting up free energy perturbation (FEP) calculations.The latest version of the Timeline plugin provides an interface for viewing temporally changing per-residue attributes of a molecular structure. It can also display temporally changing attributes of a set of VMD selections, for example a set of all the salt-bridge pairs observed in a trajectory. The controls allow selection of the molecule used for the calculation. The graphical display of residues and timesteps can be scrolled and zoomed as necessary to see results for large structures and long trajectories.The new Symmetry Tool plugin provides an easy-to-use graphical interface to the new "measure symmetry" analysis command. It determines the symmetry pointgroup of a given selection and displays the symmetry elements (inversion center, mirror planes, rotation axes, rotary reflection axes). The underlying algorithm is fairly robust and can handle molecules whose coordinates deviate to a certain extent (controlled by the tolerance parameter) from the ideal symmetry. The closest match with the highest symmetry are returned.VMD now incorporates built-in support for rendering of molecular scenes with ambient occlusion lighting using Tachyon. Ambient occlusion lighting can be used to create molecular renderings that visually highlight pockets, cavities, and crowded areas of structure, and generally look more "three dimensional". Ambient occlusion is far more effective at highlighting cavities and pockets than simpler techniques like depth cueing. VMD and Tachyon also support high dynamic range lighting (HDR) and high precision color image formats that can be selected with the use of the new "render format" text command.A newly updated tutorial is available, to guide users in experimenting with the use of Tachyon ambient occlusion lighting in their VMD renderings. Tachyon is compiled-into VMD and also supplied as a standalone program distributed with VMD. The included Tachyon ray tracer can be used to begin producing figures and movies using the advanced rendering features described above without installing any additional software.
VMD can now export volumetric coloring frequently used for display of electrostatic potentials, densities, and other spatially enumerated quantities on molecular surfaces and other space filling representations, and also frequently used for the field lines representation.Beginning with this release, VMD can add "outline" shading to the edges of any space-filling representations through the use of new material properties. Several new default materials have been added to make it easy for users to try the new shading feature in their own molecular scenes. A newly updated tutorial and example scenes show how to create VMD renderings using the new outline shading material properties. The outline material properties are also rendered by Tachyon, so they can be used in combination with ambient occlusion lighting, shadows, and other advanced rendering features.