Substrate Transport in Lactose Permease
Lactose Permease: Breaching the Barrier
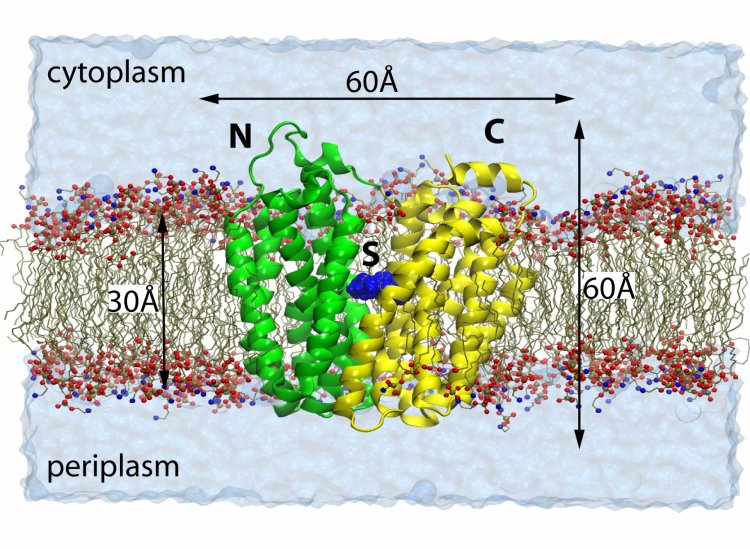 Fig. 1 Lactose
Permease embedded in a lipid bilayer.
Fig. 1 Lactose
Permease embedded in a lipid bilayer. Lactose permease (LacY) is an integral protein that facilitate the passage of lactose, one of the essential nutrients for all life forms, across the otherwise impermeable phospholipid bilayers that surround all cells and organelles. The active transport uses the energy of the electrochemical proton gradient, i.e. one H+ is transported in with each sugar (co-transport). The proteins play a critical role in transmembrane traffic, and, therefore, are critical for a healthy metabolism of a wide range of living organism, including human being. Malfunction of these transporters is associated with various pathophysiological conditions, such as diabetes and depression. Solved in 2003, the crystal structure of LacY of E. coli exhibits 12 transmembrane helices. Two halves of the protein (N-domain and C-domain, respectively) form a hydrophilic cavity opening to the cytoplasm, where the substrate is bound in its binding pocket. The periplasmic side of the protein is closed and the substrate is ready to diffuse into the cell through the opening of the cavity.
The crystal structure of LacY reprents the inward open state of the protein, in which the substrate is accessible only from the cytoplamsic side. Apparently, the accessibility of the substrate from the periplasm is necessary for the import of lactose from outside of the cell. One of the most important unknowns in the mechanism of sugar transport in LacY is the nature of protein conformational changes that switch substrate accessibility from the cytoplasmic part to the periplasmic one.
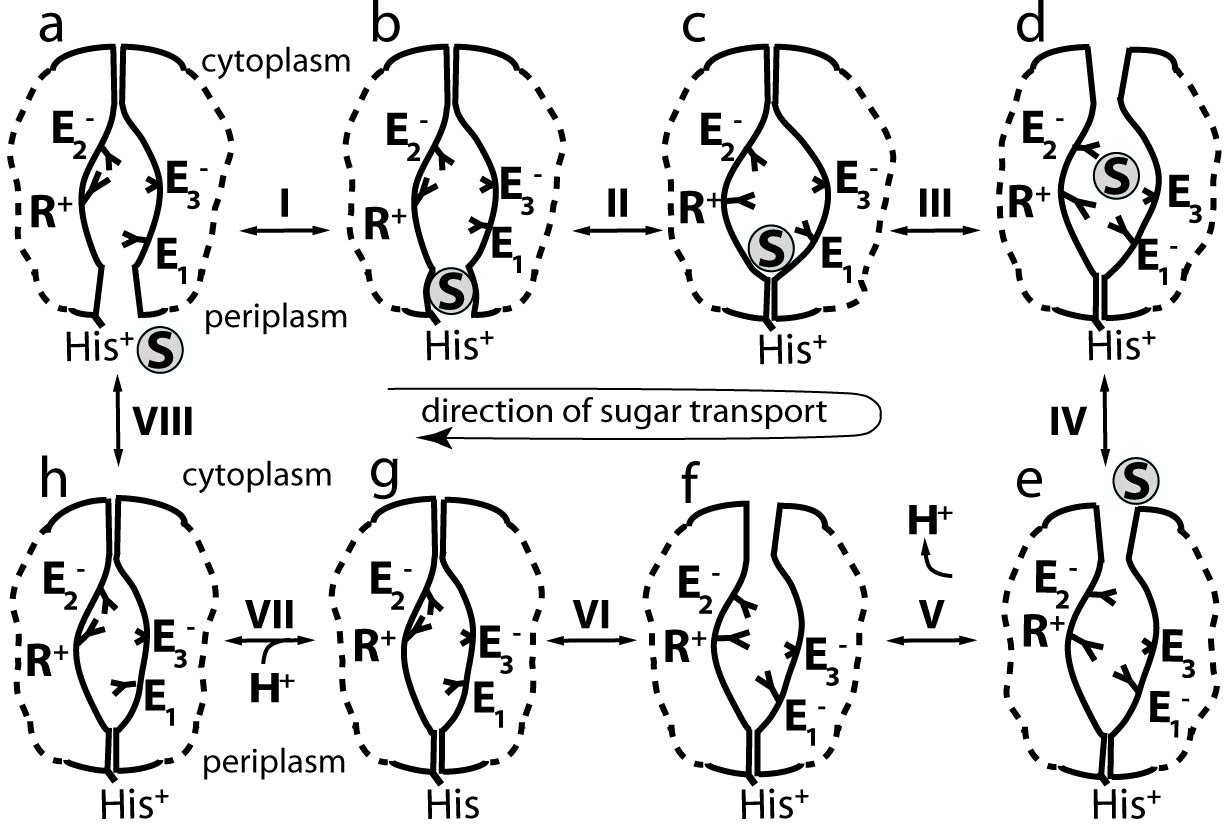
Fig. 2 Substrate transport in LacY.
Beginning from a putative outward open conformation, in which the substrate is accessible from the periplasm, the right figure provides the schematic representation of a possible lactose/H+ co-transport mechanism. Intermediates d, e, and f correspond to inward open conformation as seen in the crystal structure; intermediates a and b correspond to the outward open conformation that has not yet been resolved structurally by observation. A histidine (His in the right Figure) in the periplasmic side of LacY is postulated as a possible proton acceptor. It is protonated in the ground state (intermediate a) of LacY considering that the periplasm is rich in protons. The putative transport process shown is composed of eight steps: (I) Lactose binds to ground state LacY, in which a Glutamate E1 is protonated. (II) The periplasmic side of LacY closes, induced by the movement of the lactose to the binding pocket. (III) A proton is transfered from E1 to Glutamate E3, and Arginine R+ in the N-domain and unprotonated E1 forms a salt bridge which induces the inward open conformation d. Meanwhile lactose moves toward the cytoplasmic side. (IV) Lactose is released to the cytoplasm. (V) The proton on E3 is released to the cytoplasm; (VI) A histidine in the periplasmic half (His) transfers its proton to E1 and LacY closes at the cytoplasm. (VII) His becomes protonated. (VIII) LacY opens at the periplasm.
Protonation-Induced Transition
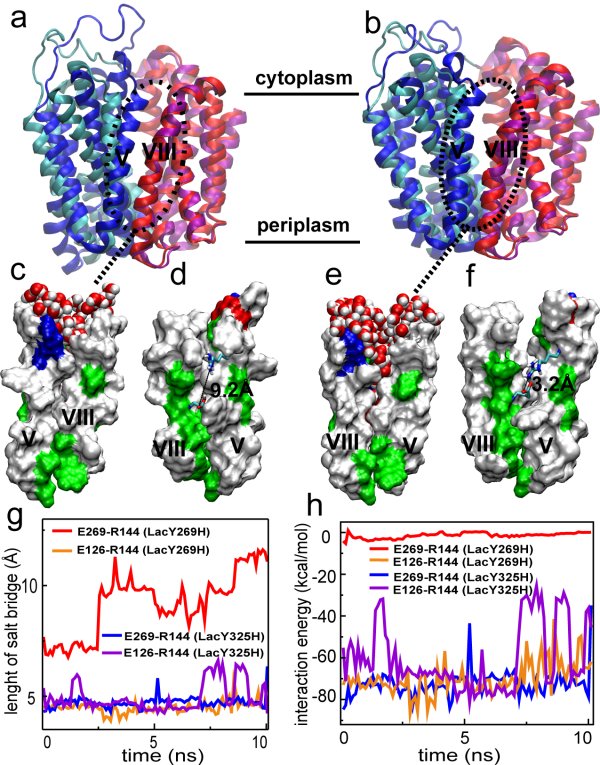
Fig. 3 Coupling of the protein conformational change and salt bridge breakage.
Our molecular dynamics simulations using
NAMD has observed the reverse of step
III, in which a proton transfer induces a conformational transition
from intermediate d to c. According to the mechanism in the above
figure, the reverse of step III includes: proton transfer from E3 to E, breakage of a salt
bridge between E1 and R+, closing of the protein
at the cytoplasm entrance and opening at the periplasmic side, as well
as substrate movement towards the periplasmic side. To mimic the effect of proton
translocation, we simulated two systems: sysa with E1
protonated and E3deprotonated, and sysb E1
deprotonated and E3 protonated. Fig. 3a and b show the
conformational changes between the crystal structure (cyan: N-domain;
pink: C-domain) and final structure (blue: N-domain; red: C-domain) of
the two systems after 10 nanosecond simulation. The
cytoplasmic halves of the N-domain helices in the final structure of
sysb show a significant inward motion
resulting in a partial closure of the cytoplasmic cavity
In contrast, no noticeable conformational changes were observed
in system sysa. In Fig. 3c,d,e, and f more details of the interaction
of the interdomain helices are shown. In sysb, the hydrophobic faces
of the interdomain helices are in a much closer contact than that in
sysa. The approach of the these hydrophobic faces appears to be controlled
by water occupancy in the interdomain region.
The breakage of the salt bridge between E1 and R+
is correlated with the conformational change of the protein.
When E1 is deprotonated, this salt bridge in sysa remain stable
during the 10 nanosecond equilibration. This same salt bridge breaks
immediately after protonation of E1 in sysb.
After the breakage of salt bridge E1-R+, R+
establishes a salt bridge with another acidic residue, glutamate E2,
which sits in the N-domain binding site.
Publications
Investigators
Related TCB Group Projects
- Mechanosensitive channels
- Structure, Dynamics, and Function of Aquaporins
- KcsA Potassium Channel
- The Dynamics of Protein Translocation
Page created and maintained by Ying Yin.



