Fibronectin and Integrin
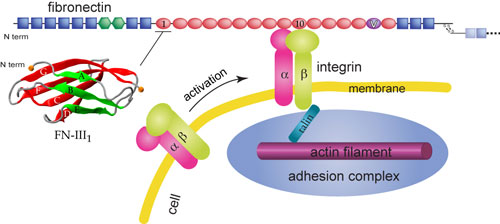
Schematic view of fibronectin bound to integrin at the cell surface.
Cells are connected through a network known as the extracellular matrix (ECM). Many cellular processes involve interactions between the ECM and the cell. The ECM not only connects cells together in tissues, but also guides their movement during wound healing and embryonic development. Furthermore, the ECM relays environmental signals to cells. One essential component of the ECM is the protein fibronectin that assembles into fibrils attaching cells to the ECM. Besides the fibrillar form, fibronectin also has a compact non-functional soluble form circulating in blood. The transformation from the compact form to the extended fibrillar form of fibronectin, a highly regulated process termed fibrillogenesis, requires application of mechanical forces generated by cells. As shown schematically on the right, cells bind and exert forces on fibronectin through transmembrane receptor proteins of the integrin family, which mechanically couple the actin cytoskeleton to the ECM via an elaborate adhesion complex.
Fibronectin
Fibronectin fibrils exhibit salient elastic properties. Cells can stretch FN fibrils up to four-fold their relaxed length. The mechanical responses of FN are conferred by its multimodular structure composed predominantly of three different repeats termed FN-I, FN-II, and FN-III. Each of the two fibronectin subunits consists of 12 FN-I, 2 FN-II, and 15 to 17 FN-III modules, respectively. While each type I or type II module contains a couple of disulfide bonds that cross-link beta-strands of the module, type III modules do not contain any disulfide bond. Structurally, FN-III modules exhibit a seven-beta-stranded sandwich motif that has been found ubiquitously in many mammalian proteins. It has been proposed that individual FN-III modules unfold to provide the elasticity of FN fibrils. In addition, stretching of FN-III modules can expose buried binding sites that serve as nucleation sites for the assembly of FN into fibrils. These buried binding sites, called cryptic sites, presumably exist either within the FN-III core or buried between the hinge regions of two neighboring FN-III modules. Cryptic sites for fibrillogenesis have been proposed to exist in FN-III1, FN-III2, FN-III7, FN-III9, FN-III10, and FN-III13-15.
Despite remarkably similar tertiary structures, FN-III modules share low sequence homology, sequence identity being typically less than 20%. Conversely, the sequence homology for the same FN-III modules across multiple species is notably higher, approximately 80%-90%, suggesting that sequence variability is functionally significant. The mechanical properties of individual FN-III modules have been compared in single molecule AFM experiments and in all-atom SMD simulation studies. The AFM experiments revealed varied mechanical stability of FN-III modules. The relative order of the stability is qualitatively consistent with the prediction obtained from SMD simulations of several FN-III modules FN-III7-10,12-14, and a FN-III module from tenascin. The simulation results have shown that FN-III modules can be pre-stretched with only minor changes to their tertiary structures before encountering the major unfolding barrier, and the mechanical stability of FN-III modules can be tuned through substitutions of just a few key amino acids by altering access of water molecules to inter-strand hydrogen bonds that break early in the unfolding pathway.
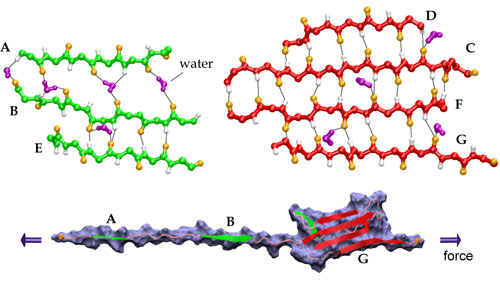
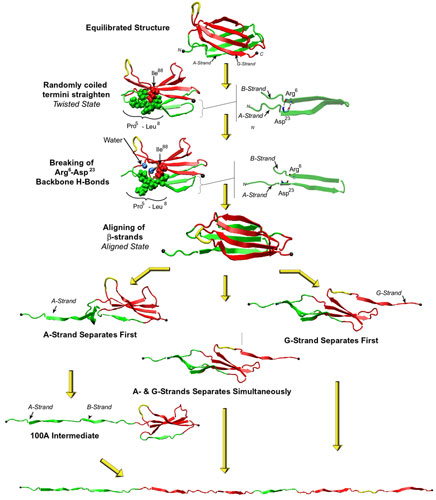
Streching a fibronectin type III module FN-III1.
Click here for a movie (MPEG, 1.7M) showing the unfolding of the module.
More interestingly, the AFM experiments with FN-III modules revealed that the mechanical response of FN-III1 is markedly different from that of other FN-III modules such as FN-III10, FN-III12, and FN-III13. FN-III1 exhibits a pronounced mechanical intermediate during forced-unfolding. While a similar intermediate has been detected for other FN-III modules such as FN-III10, it was observed less frequently than that of FN-III1. A combination of NMR structure analysis and molecular modeling provided a structural explanation of this particular property of FN-III1. Homologous to other structurally solved FN-III modules, FN-III1 consists of two four-stranded beta-sheets packed into a beta-sandwich motif. Unlike most other FN-III modules, however, FN-III1 misses in its G-strand a well conserved proline residue among FN-III modules. As a result, the key force-bearing inter-strand hydrogen bonds of the CDFG beta-sheet of FN-III1 is extended. Thus, the beta-sheet becomes much more stable than the ABE beta-sheet. Upon tension, the weaker ABE beta-sheet unravels first. The N-terminal portion of the module, the A- and B-strands, completely extend to 100 Å, three times the native length of a typical FN-III module. This new conformation amounts to a stable intermediate with the stronger beta-sheet protected by a cluster of inter-strand hydrogen bonds between G- and F-strands. The extension difference between native state and intermediate is in agreement with atomic force microscopy unfolding experiments; the intermediate structure predicted by the simulations is suprisingly close to the structure of anastellin, the 76 amino-acid N-terminual portion of FN-III1 obtained by cutting the A- and B-strands.
Fibronectin is recognized by integrins alpha5beta1 and alphaVbeta3. The primary sequence motif of fibronectin for integrin binding is a tripeptide, Arg-Gly-Asp (RGD), located on the loop connecting the force-bearing G- and F-strands of FN-III10. Multiple mechanical unfolding pathways have been observed in the SMD studies of this module because either of its two beta-sheets may become unraveled first, or the sheets may become disrupted simultaneously. In the most common pathway, the G-strand near the C-terminus detaches from the module, inducing a gradual shortening of the distance between the apex of the RGD-containing loop and the module surface, which reduces the loop's accessibility to surface-bound integrins. The shortening is followed by a straightening of the RGD-loop from a tight beta-turn into a linear conformation which implies a further decrease of the affinity and selectivity to integrins. In a second pathway, FN-III10 exhibits an intermediate similar to that observed in the simulations of FN-III1. The RGD-loop maintains its initial conformation until disruption of this intermediate. These observations suggest that the RGD-loop is strategically located to undergo conformational changes and constitute a mechanosensitive control of integrin recognition.
Besides the RGD peptide on FN-III10, FN-III9 also contains a short peptide that binds to integrin alpha5\beta1. This site, a so-called synergy site located about 32 Å from the RGD peptide, acts together with the FN-III10 binding site to enhance integrin-mediated cell adhesion. An SMD study of the FN-III9-10 tandem has shown that fibronectin-integrin binding interactions can be affected through the change of the relative distance between the two sites upon stretching. Prior to force-induced unfolding of the modules, the simulations revealed an intermediate state in which the synergy-RGD distance is increased to approximately 55 Å while both the conformations of the RGD-loop and the synergy site themselves are unperturbed. This synergy-RGD distance is too large for the RGD-loop and the synergy site to co-bind the same receptor molecule; they are thus functionally decoupled. The simulations suggest that increased alpha5\beta1-binding contributed by the synergy site, and associated downstream cell-signaling events, can be turned off mechanically by stretching the two FN-III modules.
Integrin
Naturally, the binding between integrin and fibronectin must sustain significant force in order to transmit force signals. It has been known for a long time that integrins recruit divalent cations for the ligand binding purpose. Crystal structures of integrin alphaVbeta3, available recently both in the presence and absence of a synthetic RGD ligand, provides the first atomic view of how this integrin interacts with RGD-containing ligands. The integrin acquires three divalent metal ions that coordinate the binding to the ligand. The Asp of the RGD peptide establishes a direct contact with the ion located at a site termed the metal ion dependent adhesion site (MIDAS), which is flanked by two neighboring ions located at the ligand-associated metal binding site (LIMBS) and the adjacent MIDAS (ADMIDAS).

Stretching a fibronectin-mimetic ligand from integrin alphaV/beta3.
Click here for a movie (MPEG, 2.9M)
Based on the crystal structure of the complex, computational modeling of the RGD-ligand unbinding process has produced a dynamic picture of how the alphaVbeta3 complex resists dissociation by mechanical forces. Consistent with the hypothesis that integrin-ligand binding provides a stable mechanical linkage, the unbinding requires a high force comparable to the peak force observed in the unfolding of the strongest FN-III modules. This major force peak correlates with the breaking of the contact between the Asp of the RGD-ligand and the MIDAS ion. RGD-binding to integrins does not involve a deep binding pocket that protects force-bearing contacts from attacks by free water, but rather forms a shallow crevice at the interface between the two subunits. A water molecule is tightly coordinated to the divalent MIDAS ion, thereby blocking access of free water molecules to the critical force bearing interactions. A similar scenario has been found through molecular dynamics simulations in a complex of anthrax toxin with its cell surface receptor, the primary binding site of the latter also involving a MIDAS motif.
Integrins are not constitutively active. Activation of an integrin from low ligand-binding affinity state to high ligand-binding affinity state requires conformational change that can originate from either cytoplasmic or extracellular interactions. For example, ligand binding to an integrin headpiece has been shown to induce the opening of the hinge angle between the betaA and hybrid domains in the integrin. The corresponding swing motion of the hybrid domain is linked to the switch to high binding affinity state of the integrin. The x-ray crystallographic structures of the unliganded alphaV/beta3 and the liganded alphaIIb/beta3 integrins display conformations for the closed to the open betaA/hybrid domain hinge, respectively. The movement of alpha1 and alpha7 helices of the betaA domain was observed in these two structures.
How the integrin alphaV/beta3 transitions to the high affinity conformation is debated. Although experiments link activation with the opening of the hinge angle between the betaA and hybrid domains in the ligand-binding headpiece, this hinge is closed in the liganded alphaV/beta3 integrin crystal structure. By replacing the mimetic ligand in the crystal structure of alphaV/beta3 with FN-III10, one of physiological ligands of the integrin, an all-atom model for the alphaV/beta3-FN complex was built. MD equilibration of this complex revealed that the conformational constraints of the leg domains are lifted and the betaA/hybrid hinge opens spontaneously from the initial bending conformation of the crystal structure. The simulations provides an atomic view of the allosteric pathway along which ligand-induced strain propagates via elastic distortions of the alpha1 helix to the betaA/hybrid domain hinge. The opening can be accelerated by applying external force, as discovered in SMD simulations of the complex.
Publications
-
Publications Database How the headpiece hinge angle is opened: New insights into the dynamics of integrin activation. Eileen Puklin-Faucher, Mu Gao, Klaus Schulten, and Viola Vogel. Journal of Cell Biology, 175:349-360, 2006. -
Publications Database Molecular mechanisms of cellular mechanics. Mu Gao, Marcos Sotomayor, Elizabeth Villa, Eric Lee, and Klaus Schulten. Physical Chemistry - Chemical Physics, 8:3692-3706, 2006. -
Publications Database Integrin activation in vivo and in silico. Mu Gao and Klaus Schulten. Structure, 12:2096-2098, 2004. -
Publications Database Structural insights into how the MIDAS ion stabilizes integrin binding to an RGD peptide under force. David Craig, Mu Gao, Klaus Schulten, and Viola Vogel. Structure, 12:2049-2058, 2004. -
Publications Database Tuning the mechanical stability of fibronectin type III modules through sequence variation. David Craig, Mu Gao, Klaus Schulten, and Viola Vogel. Structure, 12:21-30, 2004. -
Publications Database Structure and functional significance of mechanically unfolded fibronectin type III1 intermediates. Mu Gao, David Craig, Olivier Lequin, Iain D. Campbell, Viola Vogel, and Klaus Schulten. Proceedings of the National Academy of Sciences, USA, 100:14784-14789, 2003. -
Publications Database Identifying unfolding intermediates of FN-III10 by steered molecular dynamics. Mu Gao, David Craig, Viola Vogel, and Klaus Schulten. Journal of Molecular Biology, 323:939-950, 2002. -
Publications Database A structural model for force regulated integrin binding to fibronectin's RGD-synergy site. André Krammer, David Craig, Wendy E. Thomas, Klaus Schulten, and Viola Vogel. Matrix Biology, 21:139-147, 2002. -
Publications Database Comparison of the early stages of forced unfolding of fibronectin type III modules. David Craig, André Krammer, Klaus Schulten, and Viola Vogel. Proceedings of the National Academy of Sciences, USA, 98:5590-5595, 2001. -
Publications Database Forced unfolding of the fibronectin type III module reveals a tensile molecular recognition switch. André Krammer, Hui Lu, Barry Isralewitz, Klaus Schulten, and Viola Vogel. Proceedings of the National Academy of Sciences, USA, 96:1351-1356, 1999.
Investigators
- Mu Gao
- David Craig
- Andre Krammer
- Eileen Puklin-Faucher
- Hui Lu
- Viola Vogel
- Klaus Schulten
Related TCB Group Projects
Page created by Mu Gao.