Highlights of our Work
2025 | 2024 | 2023 | 2022 | 2021 | 2020 | 2019 | 2018 | 2017 | 2016 | 2015 | 2014 | 2013 | 2012 | 2011 | 2010 | 2009 | 2008 | 2007 | 2006 | 2005 | 2004 | 2003 | 2002 | 2001
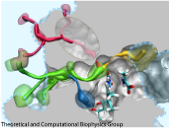
image size:
225.2KB
made with VMD
Cells contain numerous enzymes that use molecular oxygen for their reactions. Often, their active sites are buried deeply inside the protein, which raises the question whether there are specific access channels guiding oxygen to the site of catalysis. Localizing oxygen molecules in proteins is difficult. Oxygen is very mobile and difficult to locate by most experimental techniques. However, computer modeling can come to the rescue. In a recent study computational biologists using NAMD could identify the D-amino acid oxidase regions where oxygen molecules most likely reside. D-amino acid oxidase is considered essential for understanding catalysts that involve flavin moieties in proteins. The reported findings reveal a channel system through which oxygen molecules diffuse from the protein surface to the protein's catalytic center. Indeed the channel ends at a site in the protein that constitutes the ideal location for oxygen to react with flavin. The computational results are reported to be verified through so-called site directed mutagenesis experiments. Click here to read more.



