Amorphous silica
MD simulations of silicon-based nanodevices
Amorphous silica (SiO2) is an inorganic material commonly used in semiconductor circuits to isolate different conducting regions. Due to its mechanical resistance, high dielectric strength, and selectivity for chemical modification, amorphous silica has also become a key material in microelectronics and chromatography. Because of its unique properties, silica is quintessential for a broad range of applications: chips, optical fibers, and telescope glasses are manufacture on silica. Furthermore, molecular biologists employ silica in resins and optical beads to study the biomacromolecules.
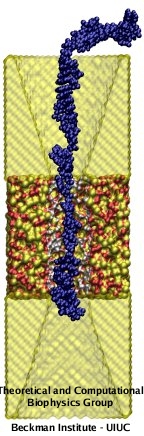
In recent years, the synergy between molecular biology and nanotechnology has opened up opportunities for many applications that involve macromolecules and silica, such as nanoelectronics, self-assembly of nanostructures, microfluidics, DNA microarray technology and nanopore sensors. The picture on the left side shows one of such nanodevices, a MOS nanopore manufactured on a poly-Silicon-Silica-Silicon membrane and a single-stranded DNA molecule (blue) translocating through it. It has been proposed that nanopores could be used to sequence DNA with single base resolution, leading to a fast and cheap DNA sequencing technology, which promises to have a enormous impact in life sciences and personal medicine. Therefore, an atomic level understanding of the interactions between biomolecules and silica is now central for further development of bionanotechnology applications.
Currently, no experimental technique is yet sensitive enough to resolve atomic-scale dynamics at the amorphous interface. Molecular dynamics simulations can be tailored to study those systems, becoming unique imaging tools. However, modeling systems that combine amorphous silica and biomolecules imposes a variety of challenges to modelers.
Until recently, computer simulations of biomacromolecules and inorganic materials, such as amorphous silica and DNA, have evolved independently from each other, and joining the expertise from both areas is a formidable task. Our previous pioneering work in nanobiotechnology was focused on crystalline inorganic structures, such as gold and silicon nitride. In this website, we briefly describe the work that the TCBG has performed to study nanosensors based of amorphous silica. For more details, take a look at the publications and links on this web site or contact the investigators listed below.
The Body: Silica Structures
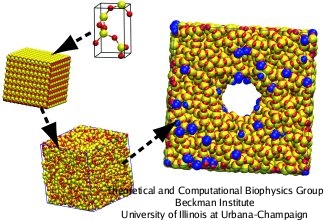
The first compulsory step to perform molecular dynamics simulations is to specify the coordinates and connectivity of each atom of the system. However, due to the amorphous nature of silica, there are no X-ray or NMR structures that can be used as templates. To obtain amorphous models, crystalline structures are randomized using a set of MD annealing steps that reproduce structural features of actual amorphous silica. The procedure to construct an amorphous silica nanodevice is squematically pictured in the figure on the right side and briefly described in the next paragraph.
First, a cristoballite unit cell is build and replicate it to fill a cubic volume big enough to contain the nanodevice. Then, the crystalline cube is annealed by increasing the temperature to 8000 K and then cooling it back slowly to 300 K. The initial high temperature assures the randomization of the crystalline structure. After that, the nanodevice is sculpted by removing atoms from the amorphous cube. To mimic real amorphous silica surfaces, the nanodevice is annealed further, allowing the rearrangement of the surface atoms. Finally, the bonding connectivity is calculated using a cutoff distance of 0.2 nm. The resulting structure can be refined to produce chemically-modified silica forms, by introducing hydroxyl groups or organic molecules at the exposed surface (blue atoms).
The detailed protocol to obtain amorphous structures is described in a paper. In that study, the annealing steps were performed for hundreds of picoseconds using the commercial software Cerius2 and specialized silica force fields (glass_2.01 and glass_2.01), which are computationally expensive. Part of our future work involves developing similar capabilities in NAMD, the premium molecular dynamics software that can run on thousand processors. Such development would allow researchers to produce amorphous models in less than a day.
The Soul: Silica Force Field
 |
 |
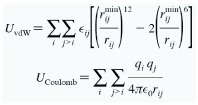
A major hurdle in simulating systems composed of inorganic materials and biological macromolecules is to describe the strenght of the interacting forces between the inorganic surface with water and biomacromolecules. A mathematical description of the system, a so-called force field, provides a set of functions for the potential energy of the system based on their atomic coordinates. We have develop a force field for amorphous silica surfaces that accurately describes the interactions with water and can be deployed to simulate nanodevices that integrate amorphous silica with biomolecules.
The hydrophobicity, a key attribute of the surface, has been used to tune silica force fields. This property can be quantified by measuring the water contact angle (WCA), the angle between the tangent of a sessile water droplet and the solid surface (see animation on the left side). The WCA value is the result of a delicate balance between specific solid interactions and the surface roughness. For silica, the WCA does not have a unique value, but depends on the amorphous topography and the concentration of exposed functional groups. The force field parameters have been refined to match the experimental water contact angles of amorphous silica. For more information about the silica parameterization and the parameters obtained, click here or refer to our paper.
�
Ion-Silica Interactions
Most applications of nanopores are based on measuring the changes in the ionic current. Therefore, understanding the ion dynamics through nanopores is desirable. Many interesting results that challenge our understanding have been reported. For instance, when immersed in electrolytic solution and under the influence of an electric field, nanopores can enhance the ion conduction for one voltage polarity but reduce it for the same voltage magnitude with opposite polarity. This phenomenon, known as ionic current rectification, has been proposed as the basis for a nanofluidic diode, a starting point for developing nanofluidic electronic devices.
There has been extensive research on the rectification phenomenon in which case the asymmetric behavior has been attributed to various factors, such as the surface charge, the ionic double layer confined in the small nanopore space and the different electrostatic distribution due to structural/chemical asymmetry at the two openings. Several models to explain rectification have been proposed based on continuous macroscopic approaches, refined to include the peculiarities of the surface system, the latter represented as a continuous surface charge. However, the applicability of continuum descriptions at the scale of a few nanometers is a subject of debate, since continuous approaches do not take into account the discreteness of the solvent nor the influence of the surface topography.
By means of molecular dynamics simulations, which provide a detailed atomistic description of the interactions between the solvated ions and the solid surface, we have performed a systematic study of the KCl conductance in silica nanopores. We found that ion-binding spots at the silica surfaces, such as dangling atoms, have effects on the ionic concentration and electrostatic potential inside the nanopore, producing the asymmetry in the I-V curve. Conversely, silica surfaces without ion-binding spots do not produce ionic current rectification. For more information about the ionic rectification, click here or refer to our recent paper.
Publications
Investigators
Related TCB Group Projects
Page created and maintained by Eduardo R. Cruz-Chu.