Atomic level structure and function of a photosynthetic vesicle
Atomic Level Structure of the Photosynthetic Unit of Purple Bacteria
Photosynthesis, the main source of energy for all life, is performed by an intricate assembly of hundreds of proteins, which harvest, transfer, convert, and store solar energy. The simplest such light harvesting machine is the purple bacterial photosynthetic unit (PSU), which performs anoxygenic photosynthesis and is significantly simpler and evolutionarily more primitive than its counterpart found in plants. The bacterial PSU is organized in the form of a pseudo-spherical membrane domain of approximately 60 nm diameter called a chromatophore vesicle.
Molecular graphics on this page
are made with VMD.
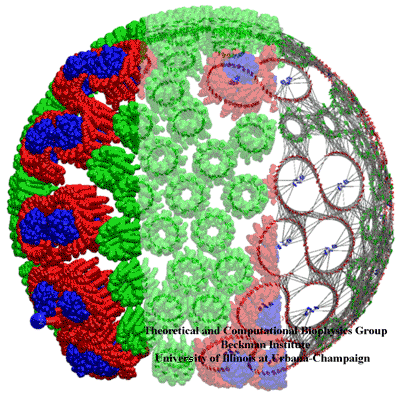
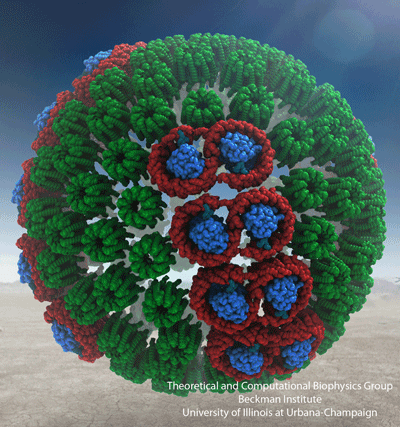
A
structural model of a chromatophore vesicle
featuring the three main light-harvesting proteins (see below).
movie (7.1 MB)
The photosynthetic apparatus of purple bacteria consists mostly of six different kinds of proteins: The light harvesting complexes, LH2 and LH1, absorb light and transfer the resulting excitation to the reaction center (RC), which subsequently initiates an electron transfer reducing quinone to hydroquinone. The LH1 and RC complexes for dimeric RC-LH1-PufX supercomplexes, where PufX is a small polypeptide likely involved in regulating the quinone traffic. The bc1 complex oxidizes hydroquinone to create a proton gradient across the membrane which in turn is utilized by ATP synthase for ATP production. Electrons are shuttled back to the RC by cytochrome c2. There are also additional proteins found in chromatophore vesicles whose structure and function are not as well understood.
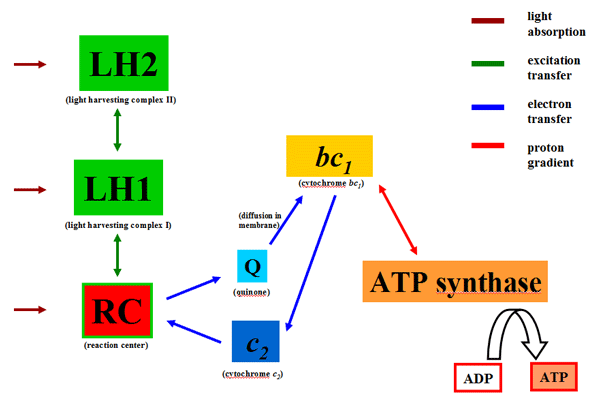
A system level view of the interactions between the
proteins that constitute the PSU.
The bc1 complexes and ATP synthase are not included in the construction depicted here as these studies focus mainly on primary excitation transfer processes. It is possible that bc1 complexes may be located within the vesicle closer to the RC-LH1-PufX complexes, even though they are not considered in this study.
Three different views of the PSU, from the supramolecular architecture
to individual chlorophylls to the energy transfer network across the vesicle.
The vesicle shown has an inner diameter of 60 nm.
Figure made with VMD.
Solving a Macromolecular Puzzle: Stitching the Vesicle Geometry from AFM, LD, X-Ray, NMR, and cryo-EM data
Scientists have determined through decades of research the three-dimensional structure of many of the individual proteins that constitute the PSU from multiple species. However, the overall architecture of the PSU remains elusive to a direct observation. Clues from spectroscopy, electron microscopy, and especially the recent AFM data of the vesicles permits us finally to reconstruct a vesicle from multiple patches computationally stitched together.
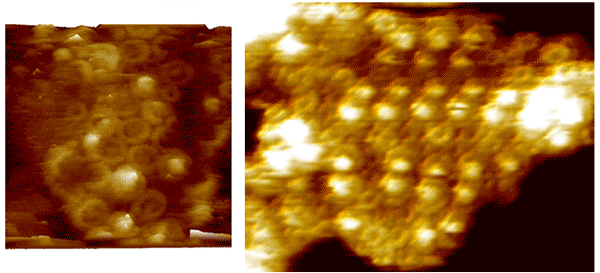
Typical AFM images used in the construction.
(Image corteousy of John Olsen.)
An area-preserving map between planar and spherical regions (depicted below) were used for the construction of small spherical patches for the protein clusters. Since both the AFM imaging procedure as well as this projection introduces deformation artifacts, resulting steric clashes and gaps were removed manually.
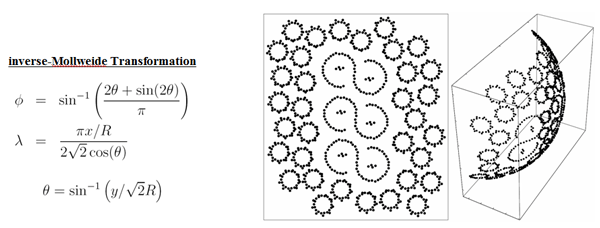
The inverse Mollweide transformation is used to map small planar patches
onto a spherical geometry before "stitching" them onto the vesicle
shown above.
Two different vesicle geometries were assembled with different LH1-RC:LH2 ratios, corresponding to different growth conditions.
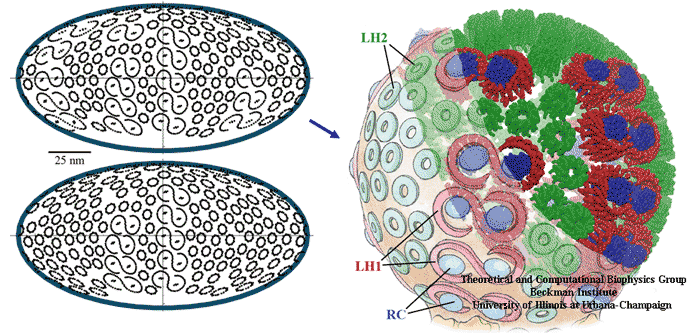
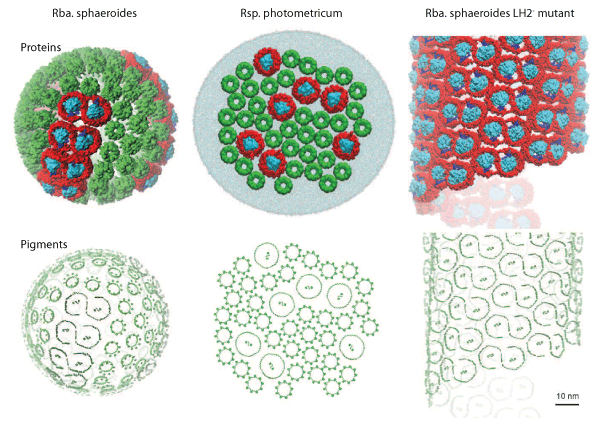
Architecture of two PSU vesicles.
Efficient Excitation Transfer across the Vesicle
The architecture of the protein assembly thus constructed enables us to model quantum mechanical excitation transfer process across the whole vesicle. The vesicle depicted has around 4000 chlorophylls belonging two over 200 clusters. This system displays a very high efficiency of 95% with an average excitation lifetime of around 50ps, which compares well with experiments. Also, a significant level of excitation sharing has been observed between neighboring reaction centers within the same dimer cluster.
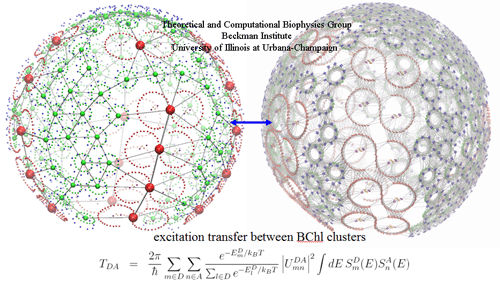
The excitation transfer networks across the vesicle, depicted
as inter-chlorophyll couplings (right) and cluster-to-cluster
transfer rates (left). The transfer between the clusters are
computed according an assumption of Boltzman equilibrium
prior to the transfer.
Figure made with VMD.
Curvature of constituent proteins and assembly of the chromatophore
There is significant variation in chromatophore shape among species. Rb. sphaeroides and Rb. capsulatus feature small vesicles; Rps. acidophila and Rs. molischianum feature flat lamellar folds; and certain Rb. sphaeroides mutants feature long tubules. This variation exists despite the main constituent proteins of the chromatophore being very similar across species. It is therefore likely that the structural variations across species are caused by the difference of the profile of curvature induced by the LH proteins in each species. Curvature properties of the chromatophore proteins has been explored via MD simulations.
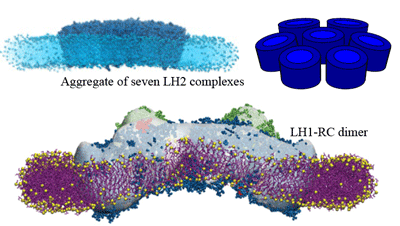
Shape of the chromatophore vesicle is determined by the
curvature profile of constituent proteins.
Top:
LH2 complexes interact
to induce membrane curvature;
bottom:
LH1-RC-PufX dimers
bend and twist the intracytoplasmic membrane.
Review Publications detailing the Photosynthetic Unit
We have reviewed the structure, function and organization of pigment-protein complexes in purple bacteria in a number of publications. In particular, how complexes affect curvature in the photosynthetic membrane is reviewd in Hsin et al., ChemPhysChem, 2010. The description of excitation transfer in purple bacteria is presented in a review aimed at undergraduates in Strümpfer et al., The light-harvesting apparatus in purple photosynthetic bacteria, introduction to a quantum biological device, World Scientific 2011. How the physics of excitation transfer is reflected in the structure and organisation of the photosynthetic unit is reviewed in Sener et al., ChemPhysChem, 2011.
Related web pages on our site
Excitation transfer in light harvestingorganization of energy transfer networks.
Quantum Biology of the Photosynthetic Unit
Light-Harvesting by Carotenoids
The Tale of Two Photosystems (on plant and cyanobacterial light-harvesting)
Related publications

(left: cover of the July 2010 issue of Biophys. J. featuring a chromatophore model .)
People
Melih Sener, Johan Strumpfer, Jen Hsin, Danielle Chandler, Chris Harrison, James Gumbart, John D. Olsen, Arvi Freiberg, Neil Hunter, Klaus Schulten.