Quantum Dynamics
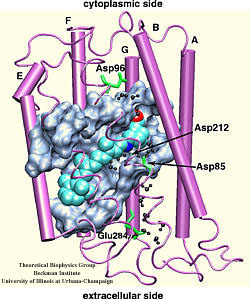
Bacteriorhodopsin (bR) is a protein that realizes the simplest known form of biological photosynthetic energy storage, absorbing light and converting its energy into a proton gradient across the cellular membrane of archaebacteria through vectorial proton translocation [1] As its name indicates, bacteriorhodopsin is closely related to rhodopsin, the protein which acts as the primary light detector in the visual pathway of higher life forms [2-6]. (click on the image for a bigger version of the picture)
In order to understand how proton transport is coupled to light absorption one must focus on the photodynamics of the retinal chromophore which intercepts the proton conduction pathway. Upon absorption of light, retinal undergoes a sub-picosecond all-trans --> 13-cis phototransformation involving torsion around a double bond. The main reaction product triggers later events in the protein that induce pumping of a proton from the cytoplasmic side to the extracellular side. The arrangement of bR is consistent with this pump mechanism. Since the primary phototransformation of bR proceeds on three coupled electronic states it can not be described via a straightforward solution of Newton's equations of motion. Instead, the quantum mechanical nature of the nuclear degrees of freedom must be confronted.
Some important stages of the bR photocycle are poorly reproduced by conventional molecular dynamics simulations due to the empirical nature of the force field employed in this method. This deficiency is particularly noticeable when processes such as photoisomerization or proton transfer are simulated. These processes require quantum mechanical evaluation of forces, since the latter are not covered by the semiempirical force field available. Due to lack of detailed quantum mechanical calculations of the retinal excited state, a particular mechanism of the initial photoisomerization step in the bR proton pump cycle still remains as an issue of controversy [5]. Some preliminary ab initio quantum mechanical calculations of the proton transfer process have been computed on model systems [6]; however, no attempt has been made to simulate this process in the real protein environment within the bacteriorhodopsin active site.
To this end, we have applied a formally exact quantum mechanical procedure, the full multiple spawning (FMS) method for molecular dynamics on multiple electronic states [7, 8]. In this first application of quantum dynamics to protein photocycles a rich set of isomerization scenarios arose with compelling suggestions regarding bR's function. In agreement with previous classical [6] and combined classical/quantum mechanical molecular dynamics studies [9], we found a sub-picosecond time scale for the key primary processes--torsion around retinal's C13=C14 bond and non-adiabatic crossings. The calculations predicted a photoisomerization quantum yield of 0.48 which is in reasonable accord with the experimentally observed quantum yield of 0.64*4 [10-13]. The simulations also revealed the occurrence of two types of photoisomerization products. These had been previously identified with a putative precursor for the pump process and with a side product which could be of possible functional significance, e.g., in case of an overcharged bacterial membrane [6, 14].
Most intriguing in regard to functional implications is the observation that photoisomerization occurs in two opposite rotational senses and that the direction of rotation is correlated with the formation of different photoproducts. It appears that one rotational sense leads predominantly to a product which is likely to trigger proton pumping whereas the other rotational sense leads to a product which has been implicated with a reversal of proton pumping as it arises in certain mutants and possibly in the presence of electric fields under intense radiation [14, 15]. The present investigation will be considerably extended in the future. Foremost is a need to improve the potential surfaces of retinal. In particular, we plan to include the effect of electronic excitation on other degrees of freedom thereby allowing for a conical-intersection. This could have a profound effect on the photodynamics since internal conversion is known to be extremely efficient when conical intersections are encountered. Another improvement that is needed is the recognition of the differing charge distributions and polarizabilities of the electronic states. This may be especially important in light of the importance of nearby ionized residues as elucidated by mutagenesis studies. A second great opportunity for further advances arises through the availability of much improved structures of bR [16, 17]. Methodological advances in modeling also permit today to simulate the integral membrane system of bR, the purple membrane, as well as to identify structral determinants of spectral tuning in retinal proteins.
Investigators
- Todd Martinez - Quantum Dynamics Simulations
- Michal Ben-Nun - Quantum Dynamics Simulations
- Ferenc Molnar - Quantum Chemistry
- Hui Lu - Molecular Dynamics Simulations
Acknowledgement
This project is funded by grants MCB-9982629 and MCB-0234938 from National Science Foundation.
Related Projects
References
- R. H. Lozier, R. A. Bogomolni and W. Stoeckenius, Biophysics Journal 15 (1975) 955.
- R. A. Mathies, S. W. Lin, J. B. Ames and W. T. Pollard, ARBBC 20 (1991) 491.
- J. K. Lanyi, J. Bioenerg. Biomemb. 24 (1992) 169
- D. Oesterhelt, J. Tittor and E. Bamberg, J. Bioenerg. Biomemb. 24 (1992) 181.
- T. Ebrey, in: Thermodynamics of Membranes, Receptors and Channels, Vol. eds. M. Jacobson (CRC Press, New York, 1993) p. 353.
- K. Schulten, W. Humphrey, I. Logunov, M. Sheves and D. Xu, IJC 35 (1995) 447.
- T. J. Martânez, M. Ben-Nun and R. D. Levine, J. Phys. Chem. 101A (1997) 6389.
- M. Ben-Nun and T. J. Martânez, JCP 108 (1998) 7244.
- W. Humphrey, H. Lu, I. Logunov, J. Herman, J. Werner and K. Schulten, (submitted) .
- G. Schneider, R. Diller and M. Stockburger, Chem. Phys. 131 (1989) 17.
- R. Govindjee, S. P. Balashov and T. J. Ebrey, Biophys. J. 58 (1990) 597.
- S. Logunov, M. El-Sayed and J. Lanyi, J. Phys. Chem. 100 (1996) 2391.
- S. Logunov, M. El-Sayed and L. Song, Biophys. J. 70 (1996) 2875.
- W. Humphrey, E. Bamberg and K. Schulten, Biophysics J. 72 (1997) 1347.
- G. Nagel, B. Kelety, B. Mockel, G. Buldt and E. Bomberg, Biophysics Journal 74 (1998) 403.
- E. Pebay-Peyroula, G. Rummel, J. Resenbusch and E. Landau, Science 277 (1997) 1676.
- Y. Kimura, Vassylyev D, M. A., A. Kidera, M. Matsushima, K. Mitsuoka, K. Murata, T. Hirai and Y. Fujiyoshi, Nature 389 (1997) 206.