Bacteriorhodopsin and the Purple Membrane
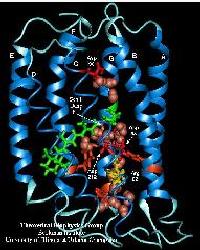
Bacteriorhodopsin (bR) is a 26~kDa transmembrane protein that acts as a light-driven proton pump in Halobacterium salinarum, converting light energy into a proton gradient. bR is the only protein constituent of the purple membrane (PM), a two-dimensional crystal lattice naturally present as part of the plasmic membrane of the bacterium. In addition to bR, PM contains ten haloarcheal lipids per protein unit that have identical saturated side chains, but differ in the nature of their polar heads. bR contains seven alpha-helices that enclose an all-trans retinal chromophore linked via a protonated Schiff base to residue Lys216. Upon light absorption, retinal undergoes an isomerization process that results under normal conditions in the translocation of a proton from the cytoplasmic side to the extracellular side of the membrane (quantum yield ~0.67). Vectorial proton translocation through membranes is a fundamental energy conversion process in biological cells. bR's utmost simplicity in comparison with other proton translocating, bioenergetic proteins makes it an ideal model for the study of this process.
Important Issues
A remarkable aspect of bR is that the minute details of its functioning are still subject of intense work and controversy, despite decades of research and impressive breakthroughs. One has today a possibility to solve many, if not all, of these problems as we have finally available high resolution structures of bR and, since very recently, of some of its photocycle intermediates on which the required physical analysis can be based. An explanation of bR's proton pumping requires necessarily an extrapolation from the known static structure and from other observational data. Proton translocation is a dynamic process and likely involves very minute motions partially below the resolution of observation, but accessible to molecular modeling. Methodological advances in modeling permit today to simulate integral membrane systems like the PM, as well as to provide highly accurate quantum chemical descriptions of potential surfaces for multiple (ground and excited) states, both preconditions for an explanation of bR's pump mechanism.
The Theoretical Biophysics Group works on three level of details of this molecular assembly:
- Structure and dynamics of the purple membrane.
- Structure and dynamics of the protein monomer.
- Quantum characterization of the retinal chromophore.
Structure and Dynamics of the Purple Membrane
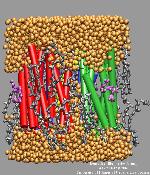
The purple membrane was built by replicating in space a hexagonal unit cell using periodic boundary conditions. The unit cell contains 3 monomeric bR molecules, 28 lipid molecules, and 8410 water molecules, giving a total number of 23783 atoms. The informations contained in several crystal structures were used to assemble all the parts of the unit cell. Click on the figure on the right to see an image (59 kB) of the unit cell's structure (figure created with VMD).
The unit cell is replicated in all directions of space, following a hexagonal symmetry, to construct the multilammelar PM model we used in our calculations. Click on the figure on the left to see an image (118 kB) of the PM layer's structure (figure created with VMD).
The crystal structures used to build the unit cell contain water molecules and external hydration was added by molecular modeling. This results in a full hydration of the unit cell (orange spheres in the picture). Click on the figure on the right to see an image (87 kB) of the hydration of the unit cell (figure created with VMD).
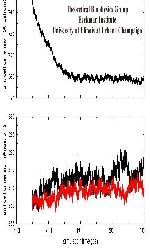
Molecular dynamics simulations of the PM model were run using the program NAMD2 to a total simulation time of 1 ns at a temperature of 300K. The last 350ps of the MD run were done in the NpT ensemble, with a constant pressure of 1 atm. The figure shos the variations of the 3 unit cell dimensions during the NpT simulation. Note the important reduction of the dimension perpendicaular to the membrane plane, and the relative constant dimensions in the membrane plane. Click on the figure on the left to see the thermodynamics of unit cell (65 kB)
.
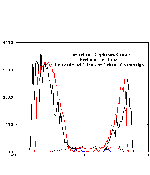
During the MD simulation, certain molecules of the unit cell undergo important rearrangements. This is the case in particular for the external water molecules (outside the protein). This picture shows the i distribution of water molecules and of the proteins' alpha carbons in the unit cell before and after the 1ns MD simulation. Note that the protein is not distribution of the protein is ~ not modified by the NpT MD, but that the water molecules are redistributed. Several water moelcules penetrate the PM in the external PGP lipid sites. Click on the figure on the right to see the variation of the PM structure after MD (10 kB) .
Publications
Acknowledgement
This material is based upon work supported by the National Science Foundation under Grant No. 0234938. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Investigators
- Shigehiko Hayashi - QM/MM and excited state calculations.
- Emad Tajkhorshid - Molecular dynamics, ab initio calculation.
- Jerome Baudry - Structural modeling, molecular dynamics simulations.
- James Phillips - Molecular dynamics, development of MD code NAMD2.