HDL and Nanodiscs
High-Density Lipoproteins (HDL)
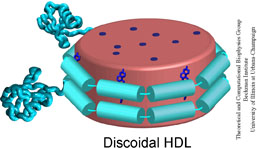
Lipoproteins are protein-lipid particles which circulate in the blood collecting cholesterol, fatty acids and lipids. One such lipoprotein particle is called high-density lipoproteins (HDL) and is often regarded as "good cholesterol" due to its ability to remove cholesterol from artery walls and transport them to the liver for degradation. Low levels of HDL have been implicated in the increased risk of coronary heart disease, the single largest killer in the United States.
The reverse cholesterol transport pathway regulates the production, transformation, and degradation of HDL particles. Apolipoprotein A-1 (apo A-1), the primary protein component of HDL, is produced in the liver and intestines. Apo A-1 initially forms a lipid-free/poor particle in which little or no lipid is associated with the protein. The efflux of lipids and cholesterol from peripheral tissues to lipid-free/poor HDL particles results in the formation of nascent discoidal HDL particles. The free cholesterol that is located on the surface is then esterfied by lecithin cholesterol acyl transferase. The esterfied cholesterol migrates to the core of the nascent discoidal HDL causing the particles to change from discoidal to spherical. These mature spherical HDL will continue to grow when additional cholesterol from cells aggregate on the surface of the particles and are subsequently esterfied. The spherical HDL are eventually degraded and recycled in the liver, thus allowing for the removal of cholesterol from tissues.
Nanodiscs
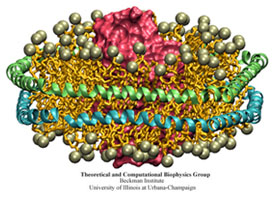
Native HDL are heterogeneous particles with a variety of shapes thus precluding structural studies of apo A-1. However, reconstituted HDL, in which purified (and often truncated) apo A-1 is used to form HDL particles with a variety of lipids, with and without cholesterol, can be made into homogeneous particles. Nanodiscs are an engineered rHDL mimic being developed by our collaborator Stephen Sligar (U. Illinois at Urbana-Champaign) which can be self-assembled using a precise set of optimized conditions to form discoidal protein-lipid particles with homogeneous size and composition. These nanodiscs are being developed as platforms in which to embed membrane proteins in a native phospholipid bilayer environment. Various membrane proteins, such as cytochrome P450s, bacteriorhodopsin, rhodopsin, translocon, blood clotting factors, and bacterial chemotaxis receptors, have already been successfully incorporated into nanodiscs. We utilized the extensive characterization of nanodiscs in furthering our understanding of the structure of apo A-1 in HDL as well as on the assembly of such particles.
Structure of Apo A-1 and Assembly of Nanodiscs and HDL
Native discoidal HDLs are heterogeneous particles which can exist in a variety of sizes. There is evidence of particles with two, three, or four apo A-1 proteins per HDL. The most widely studied of these particles are those with two proteins per HDL particle, such as nanodiscs, for which three models exist for the orientation and structure of apo A-1 around the lipid bilayer: double-belt, helical-hairpin, and picket-fence models. A variety of evidence exists for each of these models, although recent experimental and theoretical data favor either a double-belt or a helical-hairpin, but not the picket-fence model.
Lipid binding domain of apolipoprotein A-1
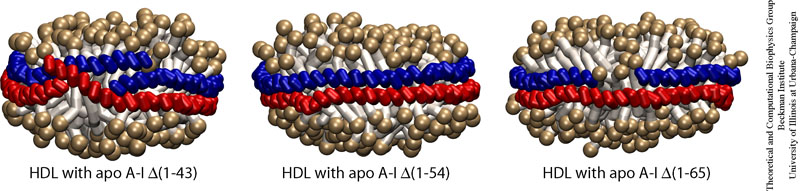
Secondary structure analysis of apo A-1 suggested a 43 residue N-terminal globular domain and a 200 residue C-terminal lipid binding domain. However, experimentally, nanodiscs prepared with truncated apo A-1 proteins (&Delta1-43, &Delta1-54, &Delta1-65), called membrane scaffold proteins, resulted in discs of identical size (9.6 nm) and composition (2 proteins and 160 DPPC lipids per disc). All-atom molecular dynamics simulations were done on these systems in order to determine the actual lipid binding domain of apo A-1 in formation of these discoidal particles (Biophysical Journal, 88:548-556, 2005). Based on mean surface area per lipid calculations, comparison of small angle x-ray scattering curves, and the relatively planar shape of nanodiscs made from truncated scaffold proteins, we concluded that the first 17 to 18 residues of the 200-residue apo A-1 lipid-binding domain are not involved in formation of the protein belts surrounding the lipid bilayer. Subsequent, coarse grained molecular dynamics simulations also confirmed the lipid binding domain of apo A-I was shorter then originally predicted (Journal of Structural Biology, 157:579-592, 2007).
Assembly of Nanodiscs & HDL
Nanodisc assembly was optimized to produce homogeneous particles. The process uses cholate (a detergent) to solubilize lipids and membrane scaffold proteins (engineered truncated apo A-1). Upon removal of cholate using either dialysis or detergent removal beads, the lipids and proteins self-assemble into discoidal particles with discrete size and composition. Due to the long time scales of such assembly processes a coarse-grained model was used to allow molecular dynamics simulations into the microsecond time scale (all-atom molecular dynamics is limited to the nanosecond time scale).
Molecular dynamics simulations of pre-assembled nanodiscs using the coarse grained
model accurately reproduced overall structural features, such as bilayer thickness and disc diameter.
Increases in temperature resulted in a decrease in bilayer thickness and an increase in disc diameter
(Journal of Physical Chemistry B,
110:3674-3684, 2006) in agreement with experimental data.
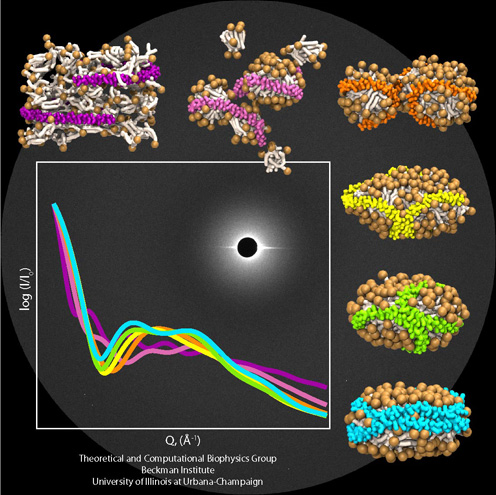
Disassembly of Nanodiscs with Cholate
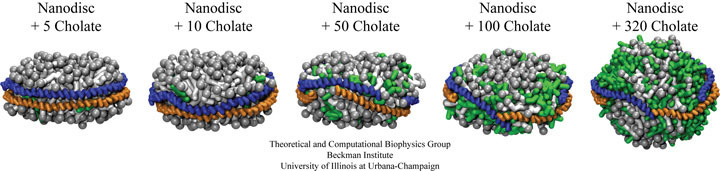
The effect of cholate on pre-formed nanodisc particles was investigated using both coarse-grained molecular dynamics and experimental small-angle X-ray scattering measurements. Small-angle X-ray scattering provides low-resolution structural information of solution state nanodiscs, allowing one to capture the "disassembly" of nanodiscs. The addition of small concentrations of cholate (5 to 10 molecules) to nanodiscs was found to have very little effect on the discoidal shape and size of nanodiscs. Additions of larger amounts of cholate (50 to 100 molecules) resulted in an opening of the scaffold proteins encircling the lipid bilayer resulting in an increase in overall particle dimension. The scaffold proteins in the coarse-grained simulations, fluctuated at the edge of the discs, never falling off the edge to lay along the top of the nanodiscs. Experimental small-angle X-ray scattering evidence suggests that given enough time, the proteins do in fact fall move from the perimeter of the lipids to lay on top of the bilayer. Addition of cholate concentrations equivalent to starting ratios used in nanodisc assembly (i.e., 320 cholate molecules), resulted in the formation of a spherical particle with a water filled interior cavity and with the scaffold proteins wrapped around the outside. This particle upon reverse coarse-graining and simulating using all-atom molecular dynamics was observed to fall apart, as would be expected (Nano Letters, 7:1692-1696, 2007).
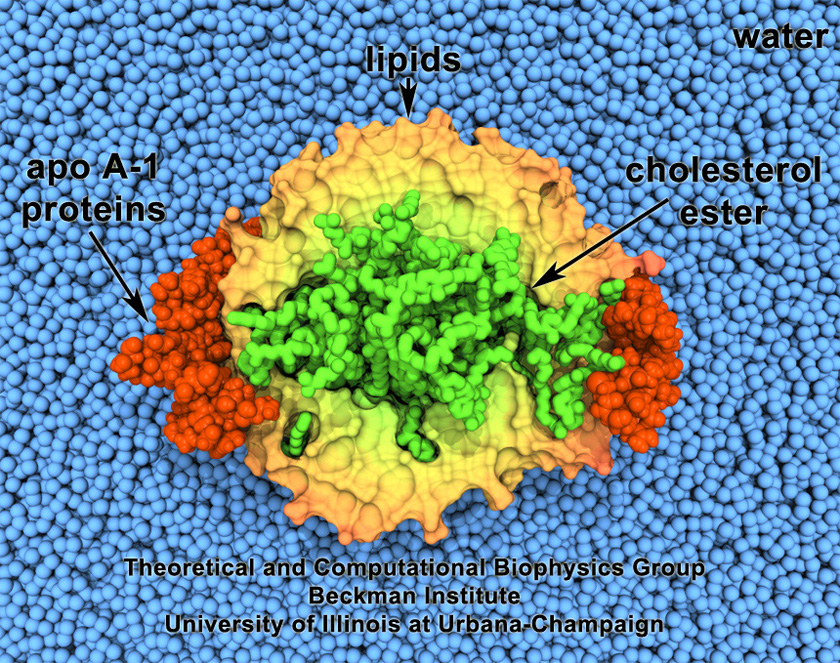
Maturation of HDL
The maturation of HDL particles from discoidal to spherical involves the conversion of cholesterol to cholesterol ester by the enzyme LCAT and the absorption of the cholesterol ester molecules. A series of molecular dynamics simulations were performed which describe the maturation of discoidal HDL into spherical HDL upon absorption of cholesterol ester as well as the resulting atomic level structure of a mature circulating spherical HDL particle. Sixty cholesterol ester molecules were added to a discoidal HDL particle containing two apolipoproteins wrapped around a 160 DPPC lipid bilayer in a stepwise fashion. The resulting matured particle, captured first in a coarse-grained description, was then described in a consistent all-atom representation and analyzed in chemical detail. The simulations show that maturation results from the formation of a hydrophobic core comprised of cholesterol ester surrounded by phospholipid and protein; the two apolipoprotein strands remain in a belt-like conformation with flexible N- and C-terminal helices and a central region stabilized by salt-bridges. The juxtaposition of the stable central region, which has been proposed to bind lectithin cholesterol acyl transferase, with the flexible terminal helices provides an ideal location for incorporating cholesterol ester into HDL particles. (Journal of the Royal Society Interface, 6:863-871, 2009).
Movies
 |
Nanodisc Assembly with Lipids Initially in a Random Configuration (mpg, 4.3M) |
Nanodisc Disassembly with 100 Cholates (mpg, 2.3M) |
Publications
Investigators
Collaborator
- Stephen G. Sligar - University of Illinois at Urbana-Champaign
Related TCB Group Projects
Page created and maintained by Amy Shih.
Last updated on August 30, 2007



