Alignment to principal axes in VMD
 The principal axes of molecules are well-defined directions that often
correspond to important symmetries. It is often desirable to
orient a molecule so that its three principal axes are aligned with
the x, y, and z directions. For example, the
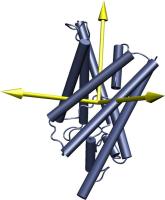
picture at right shows the water channel GlpF with its principal
axes. Because of the channel's elongated, cylindrical structure, one
axis points along the length of the channel. In order to put this
channel in a membrane, we would like to orient it so that this axis
is perpendicular to the plane of the membrane. The scripts on this
page make it easy to do this task.
The principal axes of molecules are well-defined directions that often
correspond to important symmetries. It is often desirable to
orient a molecule so that its three principal axes are aligned with
the x, y, and z directions. For example, the
picture at right shows the water channel GlpF with its principal
axes. Because of the channel's elongated, cylindrical structure, one
axis points along the length of the channel. In order to put this
channel in a membrane, we would like to orient it so that this axis
is perpendicular to the plane of the membrane. The scripts on this
page make it easy to do this task.
Installing the scripts
To use the principal axis scripts, you will need to follow these steps:
- Download
orient.tar.gzandla101psx.tar.gz. The latter is a linear algebra package by Hume Integration Software; you can look at the La homepage for more information. - Unpack the scripts
gunzip orient.tar.gz tar xf orient.tar gunzip la101psx.tar.gz tar xf la101psx.tar
- Add the following lines to your .vmdrc to make it easy to load the
packages; replace "/home/paul/scripts/" with the directory in
which you unpacked the packages:
lappend auto_path /home/paul/scripts/la1.0 lappend auto_path /home/paul/scripts/orient
Aligning a molecule to its principal axes
Load the molecule into vmd, and run the following commands to align the first, second, and third principal axes to the x, y, and z axes.
package require Orient
namespace import Orient::orient
set sel [atomselect top "all"]
set I [draw principalaxes $sel]
set A [orient $sel [lindex $I 2] {0 0 1}]
$sel move $A
set I [draw principalaxes $sel]
set A [orient $sel [lindex $I 1] {0 1 0}]
$sel move $A
set I [draw principalaxes $sel]
You may want to use the principal axes (stored in I) for
other purposes than alignment, or modify the script to get information
about the moments of inertia.



