Unfolding titin immunoglobulin domains
The giant muscle protein titin, also known as connectin, is a roughly 30,000 amino acid long filament which plays a number of important roles in muscle contraction and elasticity (Labeit et al., 1997; Maruyama, 1997; Wang et al., 1993). Titin has been connected with the diseases myasthenia gravis (Lubke et al., 1998) and hypertrophic cardiomyopathy (Rottbauer et al., 1997). The I- band region of titin, largely composed of immunoglobulin-like (Ig) domains, is believed to be responsible for the molecule's extensibility and passive elasticity. Recently accomplished AFM (Rief et al., 1997) and optical tweezers (Kellermayer et al., 1997; Tskhovrebova et al., 1997) experiments directly measured the force-extension profile of single titin molecules. In the AFM experiment, cloned sections of titin composed of adjacent I-band Ig domains were stretched at constant speed. The force-extension profile showed a sawtooth-shaped pattern with about 250 to 280 A spacing between the force peaks, with every force peak corresponding to a single Ig domain unfolding. The Ig domains were thus observed to unfold one by one, as opposed to concurrently, under the influence of applied external force Figure 1). To examine in atomic detail the dynamics and structure-function relationships of this behavior, SMD simulations of force-induced titin Ig domain unfolding were performed [1,3,7].
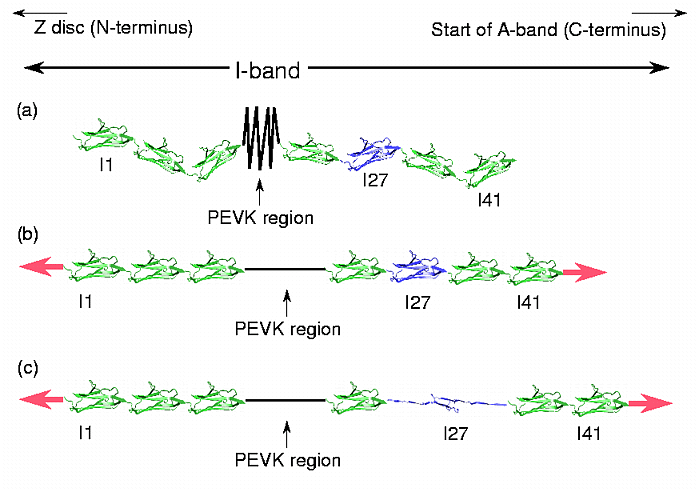
Figure 1
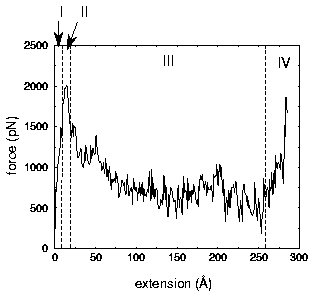
Figure 2
The SMD simulations were performed with an NMR structure of the Ig domain I27 of the cardiac titin I-band (Improta et al., 1996). I27 consists of two beta-sheets packed against each other, with each sheet containing four strands, as shown in Fig. 1b. The domain was solvated and equilibrated, then an SMD simulation was carried out by fixing one terminus of the domain and applying a force to the other in the direction from the fixed terminus to the other one. Simulations were performed following the scheme of F = K (vt - x) with v = 0.5 A/ps and 0.1 A/ps and K = 10 kB T /A2 at 300 K. The recorded force-extension profile from the SMD trajectory (see Figure 2) showed a single force peak at the initial stage of the Ig domain extension. This feature agrees well with the sawtooth-shaped force profile exhibited in the AFM experiment.
Examination of the details of the simulation trajectory provides an explanation of how the early force maximum was produced (see Figure 3). Initially (0-10 A extension), the two beta-sheets slid away from each other, each maintaining a stable structure as well as its intra-sheet backbone hydrogen bonds. As the extension of the domain reached 14 A, the structure within each sheet began to break: in one sheet, strands A' and G slid past each other, while in the other sheet, strands A and B slid past each other. The A'-G and A-B hydrogen bonds broke nearly simultaneously, producing the large initial force peak seen in Figure 2. These events marked the beginning of the Ig domain unraveling, after which the domain gradually unfolded and strands unraveled one at a time, accompanied by a large reduction in the recorded force. After an extension of 260 A, the domains were completely unfolded; further extension stretched the already extended polypeptide chain and caused the force to dramatically increase.
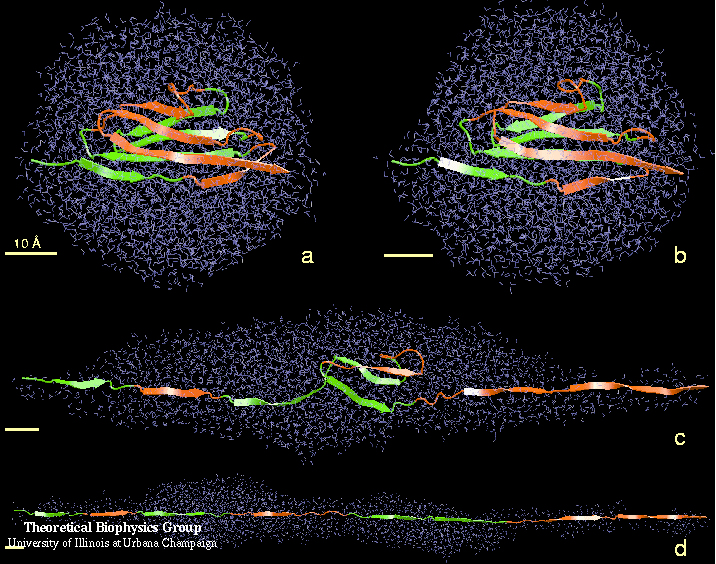
Figure 3
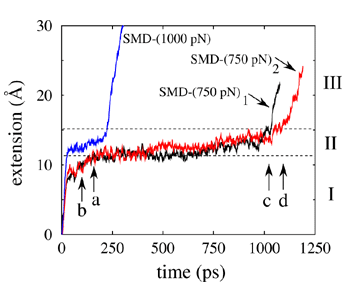
Figure 4
Constant force stretching simulations, applying 500 - 1000 pN of force, were also performed [3,4,6,7]. The resulting domain extensions are halted at an initial extension of 10 A until the set of all six hydrogen bonds connecting terminal beta-strands break simultaneously. This behavior is accounted for by a barrier separating folded and unfolded states, the shape of which is consistent with AFM and chemical denaturation data. Detailed analysis of protein water interaction shows that the breaking of hydrogen bonds between strands A' and G needs to be assisted by fluctuations of water molecules. In nanosecond simulations, water molecules are found to repeatedly interact with the protein backbone atoms, weakening individual inter-strand H-bonds until all six A'G hydrogen bonds break simultaneously under the influence of external stretching forces. Only when those bonds are broken can the generic unfolding take place, which involves hydrophobic interactions of the protein core and exerts weaker resistance against stretching [7].
The simulation suggests how Ig domains achieve their chief design requirement of bursting one by one when subjected to external forces. At small extensions, the hydrogen bonds between strands A and B and between strands A' and G prevent significant extension of a domain, i.e., the domain maintains its beta-sandwich structure. After these bonds break, resistance to unfolding becomes much smaller, and the domain unfolds rapidly. Only when a domain is fully extended does the force increase enough to begin the unfolding process in another domain.
Combining atomic force microscopy data with SMD simulation resulted in the discovery of a mechanical unfolding intermediate in titin Ig domain I27 [6]. During AFM extension of a multimer of Ig domains, before the first domain unfolding event took place, every domain was observed to extend by 6 Angstoms. SMD simulations showed that the rupture of a pair of hydrogen bonds near the amino terminus of an Ig domain (while the other inter-strand hydrogen bonds remain intact) allows this extension. Disruption of these hydrogen bonds by site-directed mutagenesis eliminates the unfolding intermediate.
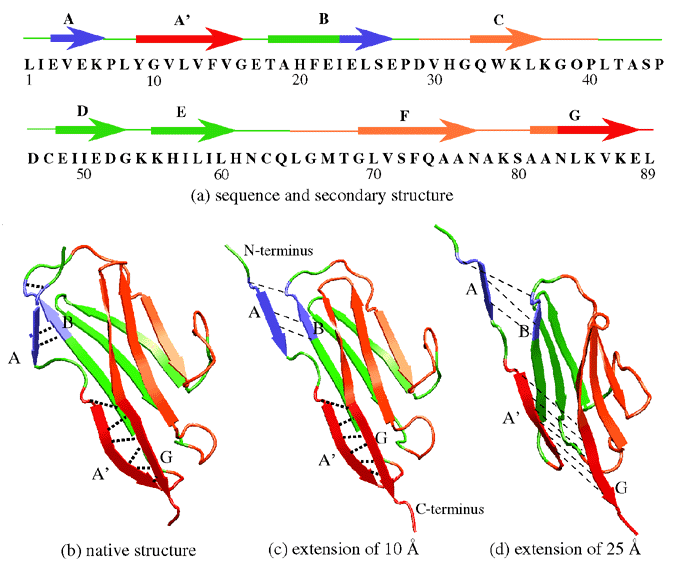
Figure 5
SMD simulation of forced unfolding of fibronectin type III (FnIII) domain (similar to the non-Ig repeated titin domains) and of other types of protein domains have also been performed [2,3]. The behavior of the proteins under external forces can be classified into 2 classes. Class I domains exhibit high resistance to forced unfolding; their fold topology are such that inter-strand hydrogen bonds must break in clusters in order to allow extension of the domain. This class includes titin Ig and FnIII domains. Class II domains, for example all-helix domains, have topologies that can be extended while breaking inter-strand hydrogen bonds singly, they do not exhibit dominant force peaks when stretched in SMD simulations.
Animations
Investigators
- Hui Lu
- Barry Isralewitz
- Mu Gao
- Andre Krammer, University of Washington
- Viola Vogel, University of Washington
- Julio Fernandez, Mayo Clinic
Publications
References
- Improta, S., Politou, A., and Pastore, A.: Immunoglobulin-like modules from titin I-band: extensible components of muscle elasticity. Structure. 4 (1996) 323-337.
- Kellermayer, M., Smith, S., Granzier, H., and Bustamante, C.: Folding-unfolding transition in single titin modules characterized with laser tweezers. Science. 276 (1997) 1112-1116.
- Labeit, S., Kolmerer, B., and Linke, W.: The giant protein titin: emerging roles in physiology and pathophysiology. Circulation Research. 80 (1997) 290-294.
- Lu, H., Isralewitz, B., Krammer, A., Vogel, V., and Schulten, K.: Unfolding of titin immunoglobulin domains by steered molecular dynamics simulation. Biophys. J. Submitted.
- Lubke, E., Freiburg, A., Skeie, G. O., Kolmerer, B., Labeit, S., Aarli, J., N.E. Gilhus, R. W., Wussling, M., Ruegg, J., and Linke, W.: Striational autoantibodies in myasthenia gravis patients recognize I-band titin epitopes. Journal of Neuroimmunology. 81 (1998) 98-108.
- Maruyama, K.: Connectin/titin, a giant elastic protein of muscle. FASEB J. 11 (1997) 341-345.
- Rief, M., Gautel, M., Oesterhelt, F., Fernandez, J. M., and Gaub, H. E.: Reversible unfolding of individual titin immunoglobulin domains by AFM. Science. 276 (1997) 1109-1112.
- Rottbauer, W., Gautel, M., Zehelein, J., Labeit, S., Franz, W., Fischer, C., Vollrath, B., Mall, G., Dietz, R., Kubler, W., and Katus, H.: Novel splice donor site mutation in the cardiac myosin-binding protein-C gene in familial hypertrophic cardiomyopathy. Journal of Clinical Investigation. 100 (1997) 475-482.
- Tskhovrebova, L., Trinick, J., Sleep, J., and Simmons, R.: Elasticity and unfolding of single molecules of the giant protein titin. Nature. 387 (1997) 308-312.
- Wang, K., McCarter, R., Wright, J., Beverly, J., and Ramirez-Mitchell, R.: Viscoelasticity of the sarcomere matrix of skeletal muscles. Biophys. J. 64 (1993) 1161-1177.



