Molecular Dynamics Study of Rhodopsin
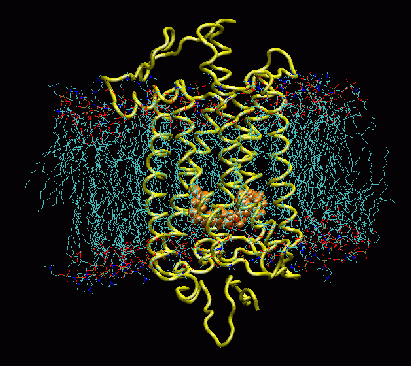 Rhodopsin embedded in a patch of POPC membrane. Retinal is shown in orange
Rhodopsin embedded in a patch of POPC membrane. Retinal is shown in orange
We have two modes of vision faciltated by two different kinds of cells in our retina. The so called cones enable color vision. Three subtypes can be distinguished which are responsible for absorption of blue, green and red light respectively. In twilight we use the rod cells which are much more sensitive to low light intensities but cannot distinguish colors.
At the beginning of the visual process is the 7-transmembrane protein rhodopsin. It belongs to the family of G-protein-coupled receptors (GPCRs) which switch from an inactive to an active form after the binding of a ligand. The activated receptor can trigger the hydrolysis G-protein bound GTP which is the beginning of an intracellular signal cascade. GPCRs are responsible for the transduction of a great variety of signals including hormones and neurotransmitters.
In case of rhodopsin the ligand retinal is covalently bound to receptor and works an inverse agonist locking the protein in the quiescent state. The change of the chromophore structure upon cis-trans photoisomerization triggers the transition of the receptor into the active form which activates the G-protein transducin. The system is regenerated by release of the trans-chromophore and binding of a new one in cis-conformation. The complete cycle takes place on the order on milliseconds but the photoisomerization itself is with 200fs among the fastest photochemical reactions.
We have studied the effects of the cis-trans isomerization of retinal on the conformation of the protein using molecular dynamics simulations. For this purpose a model of bovine rhodopsin based on the crystal structure determined by Palczewski el al. [1],[2] was embedded in a patch of palmitoyl-oleoyl-phosphatidylcholine (POPC) membrane and solvated it in water which results in a total of about 40000 atoms. Retinal isomerization was induced by switching the dihedral potential and charges of retinal to those in the excited state. After isomerization, the system was simulated for 10 nanoseconds in order to explore the effect of isomerization on the structure and dynamics of rhodopsin Saam et al, 2002.
References
- K. Palczewski et al., Science 289, 739 (2000)
- D. C. Teller, T. Okada C. A. Behnke, K. Palczewski, R. E. Stenkamp, Biochemistry 40, 776 (2001)



