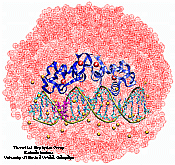
DNA Binding Domain
|
The goal of our study of the estrogen receptor interacting with specific and
non-specifis sequences of DNA was to understand and explain what effect binding
of ER has on the conformation of the target DNA, and how differences in the DNA
sequence affect ER binding specificity
To answer these questions two systems were created:
The results of our study show that the binding specificity of the ER-DBD to different sequences of DNA is correlated with how the side--chains of the protein position themselves relative to the bases of the DNA, most important being whether the side-chains can be accommodated without forming negative interactions with the bases. Also, while some hydrogen bonds are broken by the side-chains rearrangement in the non-consensus ER-G/ERE system, other bonds are created by the inclusion of water molecules. There is still an energy difference between the consensus and non-consensus complexes and that may arise from the less favorable geometry of the new hydrogen bonds. The water molecules found at the protein-DNA interface act favorably by filling gaps resulting from imperfect complementarity between interfaces and they also extend the interaction of some of the residues beyond their direct contact. Our simulations also demonstrate that interaction of the receptor dimer with DNA results in a bent and underwound conformation of the DNA compared to the initial crystallographic structure. The bend is described as a displacement or ``jog'' in the path of the helical axis when projected into a plane. We suggest that the induced bend in the DNA is a general effect of a hormone receptor dimer binding to DNA and that the bend reflects the symmetry of the dimer. The binding of the estrogen receptor dimer to the DNA and the conformational changes induced in the axis of the DNA during the simulations are presented in an animated gif movie. Detailed results can be found in this poster.
Publications
References |