Elastic Rod Model of Lactose Operon DNA
Lactose operon repressor (lac repressor) is a protein that regulates lactose consumption by E.coli bacteria. At the dawn of molecular biology, it was the study of this protein that inspired the concept of protein synthesis control via repression of gene transcription. The lac repressor binds DNA in the absence of lactose, and prevents RNA polymerase from beginning the transcription. Thus, the cell does not produce the proteins responsible for lactose digestion when there is no need for them.
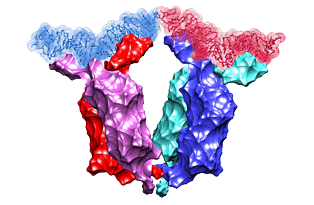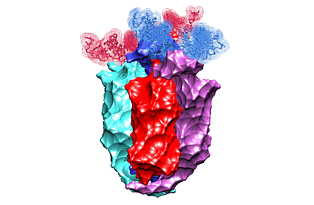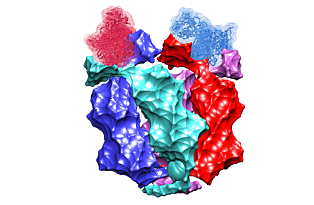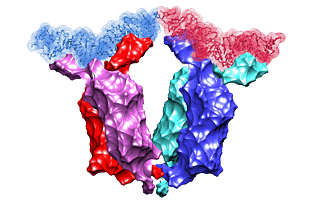
Lac repressor functions as a tetramer. Two pairs of sub-units bind two out of three DNA sites recognized by the protein. The binding induces looping of the DNA connecting the binding sites. In 1996, the crystal structure of the lac repressor-DNA complex was reported. The protein tetramer was binding two synthetic DNA segments, emulating the natural binding sites. However, those were disjoint segments not connected with an intermediate DNA. Such an intermediate DNA segment would introduce a certain stress that would possibly alter the complex structure. We use the crystal structure to fill the gap, to predict the shape of the connecting DNA loop, and to estimate the resulting stress.
We approximate the DNA in the loop by an inextensible elastic rod. The elastic rod model describes the DNA in terms of its center-line contour, the curvatures in two principal directions of the DNA cross-section, and the twist around the helical axis. The parameters of the model are two DNA bending moduli, which can be estimated from experimental data. The theory of elasticity results in a 13th order Kirchhoff system of differential equations, which completely describes the elastic rod. The crystal structure provides boundary conditions for the problem. We use and a continuation method to numerically solve the resulting boundary value problem. The method consists in gradual changes of the elastic moduli and boundary conditions, from the values corresponding to a known solution to the desired ones.