Detecting DNA Orientation with a Nanopore
In a remote New Mexico dessert, Klaus Schulten met Amit Meller, who told him a story of two DNA hairpins. The first one, when threaded through the transmembrane pore of alpha-hemolysin blocks the ionic current to 12 pA and escapes from the pore about three times slower than the other one that blocks the current to only 9 pA. The amazing fact about these experiments was that the two DNA hairpins were identical in sequence. The only difference was in the global orientation of the single stranded part of the hairpin in the pore of alpha-hemolysin. Astonished by the outcome of their experiments, Amit Meller and Jérôme Mathé sought an explanation from the modelers (Klaus Schulten and Aleksei Aksimentiev). Cautious as the experimentalists are, they did not tell the modelers which of the DNA orientations produces the larger current blockade and which one escapes faster from the pore. Below we describe our quest for the solution of the puzzle. A formal report can be found here.
Deoxyribonucleic acid (DNA)
 Double stranded DNA
Double stranded DNA
 Single stranded DNA
Single stranded DNADNA direction. Yes, a DNA strand has a direction. It is determined by the arrangement of the phosphate and deoxyribose sugar groups along the DNA backbone. One of the DNA ends terminates with the O3-H group (the 3' end), whereas the other one terminates with the O5'-H group (the 5' end). Note that there are N-1 phosphate groups in an N-nucleotide ssDNA. All sequences of DNA are usually written from 5' to 3' end. When forming a double helix, the complimentary DNA strands are oriented in opposite directions.
The alignment of the bases can indicate the global orientation of a DNA strand
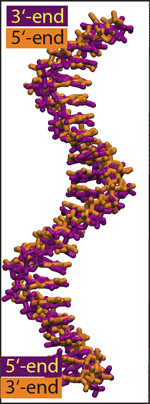 Alignment of two
poly(dA20) strands that differ from each other only by
their global orientations.
Alignment of two
poly(dA20) strands that differ from each other only by
their global orientations.
DNA strands are hard to destinguish in solution.
To reveal how the global orientation of a DNA strand determines its escape time from a nanopore, as well as the blockade of ionic current, we examined in atomic detail conformations of DNA in solution and inside alpha-hemolysin. Before running MD simulations, two identical poly(dA20) strands, different only by their global orientation, were aligned with each other, as shown in the right figure. The two strands appeared to have, overall, similar conformations, the only difference visible being the ordering of the backbone atoms: When moving along the strand from 5 to 3 end, a phosphate group follows the deoxyribose sugar ring, the C5 carbon of deoxyribose follows the phosphate. Reversing the order changes the orientation of deoxyribose in the backbone. Although this change is sufficient to disrupt the function of a dephosphorylative enzyme, when put into the perspective of the alpha-hemolysin structure, it is an unlikely reason for the observed directionality.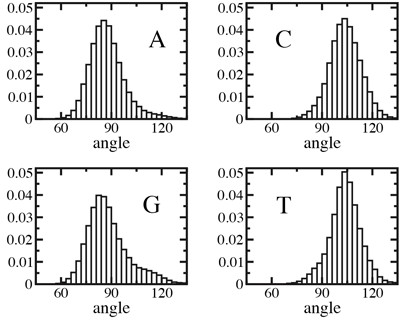 Normalized distributions of the angle between the base and the backbone of A, C, G, and T nucleotides in an unrestrained DNA strand. The average was taken over a 12.5-ns molecular dynamics trajectory.
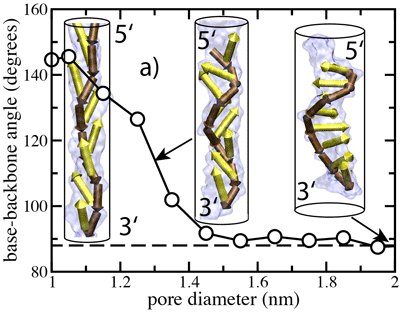
Normalized distributions of the angle between the base and the backbone of A, C, G, and T nucleotides in an unrestrained DNA strand. The average was taken over a 12.5-ns molecular dynamics trajectory.
Another possible explanation for the observed 3'- and 5'- directionality might be that an ensemble of DNA conformations, by itself, contains the information about the global orientation of a DNA strand. To determine whether this is the case, we computed the average angle between nucleotide bases and the backbone in a DNA strand submerged in water. Our results shown at the left indicate that for purine nucleotides (A and G) the most probable angle is ~88°, whereas for pyrimidines (C and T) that angle is ~105°. From this analysis, we conclude that local conformations can indeed indicate a global orientation preference of a DNA strand, although the conformational anisotropy in an unrestrained strand is small and depends on sequence.
Stretching DNA tilts its bases.
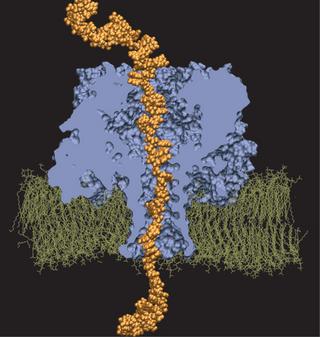
360,000-atom MD simulation reveals the mechanism of the direction-specific interaction between DNA and alpha-hemolysin
 Molecular dynamics simulation of DNA permeation through alpha-hemolysin.
Molecular dynamics simulation of DNA permeation through alpha-hemolysin.
What consequence does the uniform tilting of the bases have on DNA translocation kinetics and on current blockades? Intuitively, one could equate driving ssDNA with an electric field through the narrow constriction of alpha-hemolysin to bringing a tree through a door: The tree moves more easily trunk first than tip first (because branches tend to grow toward the tip of the tree). Naturally, to determine the molecular mechanism underlying DNA directionality, one has to visualize the translocation event in atomic detail. Currently available computer resources only allow MD simulations of a few tens of nanoseconds. However, one can start a simulation with the DNA strand threaded halfway through the pore and inspect over a few nanoseconds in how far switching on an electrical field drives the strand for the next few nanoseconds through the pore.
We have adopted this procedure to test the directionality dependence of the field-induced transport of ssDNA. Two systems, each comprising one alpha-hemolysin channel, a DPPC membrane, a poly(dA58) strand, water, and ions, were subject to an external electric field equivalent to a 1.2-V bias. In the first system, the 5' end of the strand was located at the trans side of the membrane; the system is shown in the figure. In the second system, the 5' end was located at the cis side. The applied bias forced DNA strands toward the trans compartment.
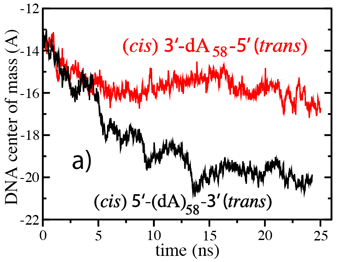
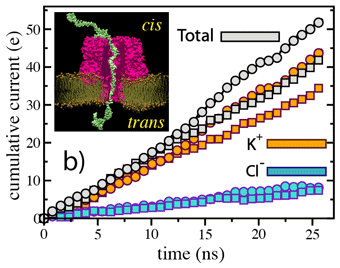 (cis)5-dA58-3(trans) data are
plotted as squares; (cis)3-dA58-5(trans) data are plotted as
circles.
(cis)5-dA58-3(trans) data are
plotted as squares; (cis)3-dA58-5(trans) data are plotted as
circles.
In conclusion, through large-scale molecular dynamics simulations (and a lot of creative thinking) we were able to reproduce the experiments results in silico and uncover the molecular mechanism of the observed directionality. To our delight, simulations and experiments converged!
Publications
Investigators
Simulations ExperimentRelated TCB Group Projects
Page created and maintained by Aleksei Aksimentiev.



