Torque Application to ATP Synthase Central Stalk
Adenosine triphosphate (ATP) is the fuel of life; every living cell must use ATP to carry out its functions. The human body synthesizes its own weight in ATP every day. This synthesis is performed by ATP synthase, a complex of two joined rotating molecular motors. A detailed understanding of how this system functions can impact areas ranging from neurodegenerative disease research to nanotechnology development. First, sugars are metabolized to crowd hydrogen atoms on one side of a membrane. Second, the hydrogen atoms push back across the membrane through a large protein complex called ATP synthase. The flow of hydrogen atoms drives two joined rotating mechanisms in each ATP synthase, resulting in synthesis of ATP from surrounding chemicals. We use MD simulations to study the rotary mechanism which converts torque into cyclical changes of ATP synthase's shape.
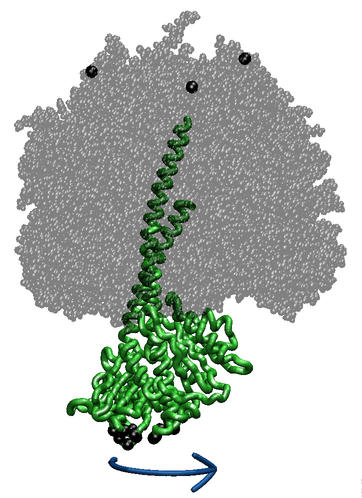
ATP synthase is a large (about 100,000 atom) multi-protein complex which includes a transmembrane Fo unit coupled to a solvent-exposed F1 unit via a central stalk. The Fo unit converts a transmembrane electrochemical potential (proton-motive force) into mechanical energy used to rotate the F1 central stalk. The stalk rotates within the surrounding subunits of F1, leading to cyclic conformational changes in the three catalytic sites in F1 and, thereby, to ATP synthesis. ATP synthase can also function in the reverse direction, hydrolyzing ATP and utilizing the released energy to pump protons across the membrane.
The three ATP synthase catalytic sites are seen in X-ray structures of F1, one in each of the subunit pairs arranged pseudo-symmetrically around the central stalk. The Fo-driven rotation of the F1 central stalk relative to the surrounding subunits is thought to drive the sites through at least three conformational states per rotation, with each site synthesizing one ATP molecule per rotation. The reverse reaction has been directly visualized; when ATP is hydrolyzed by F1, the central stalk is observed to spin, with the cyclic changes in F1 presumably producing the required torque. Conformation changes in the surrounding pairs also appear to be cooperative, with the coupling likely being transmitted both between neighboring pairs and through the central stalk. Mutation experiments have identified residues important to pairs-stalk interactions and several schemes for F1 synthase subunit interactions have been proposed.
Our simulations examine in detail how the interaction and deformation of the F1 subunits couple an applied torque to the cyclic transformation of the catalytic binding sites. There is a large body of experimental investigation into the interactions among these subunits and how these relate to F1 function. Results range from estimates of the elastic moduli of the central stalk to identification of specific residues likely involved in inter-subunit force transmission. However, many mechanical details and properties of the rotating system, which could help explain torque--catalysis coupling, can not be obtained experimentally.
We use steered molecular dynamics (SMD) simulations to apply a torque to F1, in order to directly examine the cooperative interactions between all parts of F1 and observe how the conformations and energies in our model evolve as a torqued rotation progresses. This allows us to characterize: 1) the function of the coupling mechanism, including residues involved in inter-subunit communication, large conformational changes during the cycle, and roles of elastic energy storage throughout the cycle steps; 2) the state of the catalytic sites during torqued rotation, including residues and water molecules involved in stabilizing transition states, and nucleotide binding/release.



