DBP8: Bacteria and Eukaryal Systems
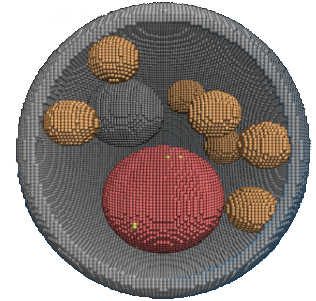
Hematopoietic stem cells (HSC) are responsible for maintaining the body's blood and immune cells; dysregulation of their behavior can lead to severe pathologies like leukemia. Understanding their proliferation and diffentiation could not only shed light on pathological processes, but could also help to expand HSC populations for transplantations. However, such understanding requires the study of gene expression and regulation, of signaling within the highly compartmentalized eukaryotic intracellular space, and of cellular interactions with complex environments - processes that can be investigated in simpler model systems. Here, we will study (1) LeuRS:tRNA-mediated intercellular signaling in Escherichia coli (S1) as a model for signaling by cytokines, especially stem cell factor (SCF), (2) the spliceosome assembly in Saccharomyces cerevisiae (S2) as a model for multi-compartmental cellular processes, and (3) HSC proliferation and differentiation in an in vitro niche environment (S3).
The first challenge arises from the sheer system size of eukaryotic cell models (S2, S3), surpassing even the largest reaction diffusion master equation (RDME) simulations performed in Lattice Microbes (LM) thus far and requiring the new multi-level parallelisation paradigm provided by MLP-LM (TRD3). The second challenge is presented by the complexities of the modeled systems in S1-3, which feature higher numbers of species and reactions as well as higher densities of particles than could be handled in LM before; this is solved by removing these limits with BIG-LM (TRD3). The third challenge lies in coupling different computational representations for different model components in S1-3: stochastic processes like gene expression, high-concentration intra- and extracellular metabolites and effectors with or without spatial resolution and, for multi-cellular systems, steady-state descriptions of metabolism. Combining these different representations in \LM will be enabled with Multiscale-LM (TRD3). The fourth challenge arises from the high degree of compartmentalization and spatial complexity in eukaryotic cells featured in S2-3, the modeling of which requires the integration of several different types of experimental data like cryo-ET, single-cell super-resolution imaging and -omics. An effective platform for this will be provided by density map segmentation and manipulation (TRD2) tools and Exp-2-LM (TRD3). The fifth challenge arises from the need to inspect and analyze the complex cellular models for S1-3. To this end, enhancements for cellular visualization and analysis (TRD2) in \VMD will provide visualization semantics for the numerous components of cellular RDME models.
This DBP covers a range of objectives from bacterial via eukaryal model systems up to a detailed whole-cell model of HSC proliferation. To fulfill these objectives, it builds on the Center's extensive experience and success in modeling cellular systems: From stochastic and reaction diffusion models of intracellular processes to systems-level descriptions of cell populations and spatially resolved multi-scale models of colonies. On the other hand, it draws on our longstanding collaborations with experts from across the imaging community, as well as our new collaboration with an expert in stem cell biology and HSC niche design. A newly discovered form of intercellular E. coli signaling, involving the cleaving and excretion of fragments of LeuRS:tRNA, will be modeled using experimental data on LeuRS secretion (Martinis). Coming to the first eukaryal model system, a complex, multicompartmental reaction diffusion model of the spliceosome assembly in S. cerevisiae will be constructed incorporating cryo-ET data and in vivo imaging data (Villa, Gruebele). These preceding model systems will set the stage for comprehensive, multi-scale models of HSC populations in different niche environments. Specifically, we seek to explain the experimental finding that HSCs react differently to the cytokine stem cell factor (SCF) in their environment when it is freely diffusing versus when it is immobilized. To study this effect, we will construct a model of SCF receptor binding, internalization (for the freely diffusing case), potential biomechanical signals that may stem from interactions with an immobilized SCF, and the downstream regulatory cascades that ultimately lead to cell differentiation. This endeavor will require experimental information on HSC growth and behavior in different engineered environments (Harley) as well as cryo-ET data on the cell interior in HSCs (Baumeister).
