Bacterial Chemotaxis
Topics: Introduction - The Chemosensory Array - CheY Activation - Publications - Additional resources - Investigators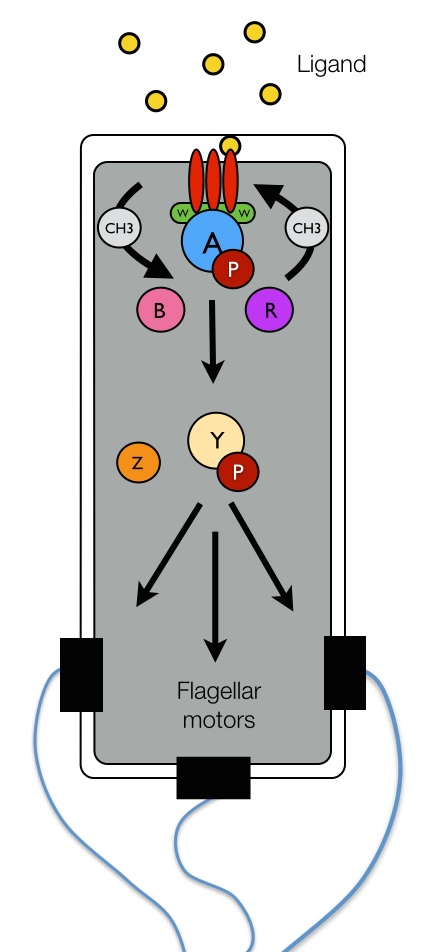
Figure 1: E. coli chemotaxis network.
Introduction
Bacterial Chemotaxis
To Tumble Or Not To Tumble? Chemotaxis is a fundamental sensory phenomenon by which biological cells translate environmental chemical information into motile behavior. Bacteria, in particular, use chemotaxis to position themselves within the optimal portion of their habitats by monitoring the environmental concentration gradients of specific chemical attractant and repellent ligands (termed chemoeffectors). In Escherichia coli the motile behavior is reduced to a binary response: change direction (tumble) or keep going (run). Through a series of interspersed run and tumble maneuvers, biased by chemical cues, cells are able to move on average towards favorable growing conditions while avoiding unfavorable ones. How do cells decide whether to run or tumble and when to do so?
Sophisticated protein networks allow bacteria to couple the physical binding of chemoeffector ligands to the rotational bias of their flagellar motors and ultimately the cell's swimming pattern. The chemotactic network of E. coli, pictured schematically in Figure 1, has served as a paradigm for the general study of bacterial chemotaxis. Environmental information is obtained by the network through specific protein receptors, termed chemoreceptors (red), on the surface of the bacterium's inner membrane. The binding of chemoeffector ligand to a chemoreceptor leads to the transduction of a signal across the cell membrane, which is used to regulate the autophosphorylation activity of the CheA kinase (blue) with the help of an adaptor protein CheW (green). The activity state of CheA is read by another protein CheY (yellow) that accepts a phosphate group (maroon) from activated CheA. Phosphorylated CheY diffuses to the flagellar motors where it binds to shift the rotational bias of a motor from counter-clockwise (CCW) to clockwise (CW). CW rotation of the flagellar motors gives rise to tumbling, while CCW motor rotation elicits runs.In addition to the protein components described above, which are universal among chemotactic bacteria, several other proteins assist with the mechanics of signal regulation in E. coli. In particular, CheZ (orange) removes phosphate groups from CheY, suppressing the chemical connection between activated CheA and the flagellar motors and terminating the tumbling bias induced by phosphorylated CheY. Two other enzymes, CheR (purple) and CheB (pink), endow cells with the ability to tune their chemotactic sensitivity to stimulus intensity in a process termed adaptation. Through the reversible methylation of several specific amino acid sites on the chemoreceptors, these two enzymes either enhance or hinder the ability of chemoreceptors to regulate the activity of CheA. Importantly, due to relatively slow rate of receptor methylation (compared to ligand binding), the methylation levels of the chemoreceptors provide a record the the cell's immediate chemical past, giving rise to short-term molecular memory. Remarkably, the cell can use this record to compare its past chemical environment with the one currently encoded in the receptor occupancy and decide what to do next.
The Bacterial Brain
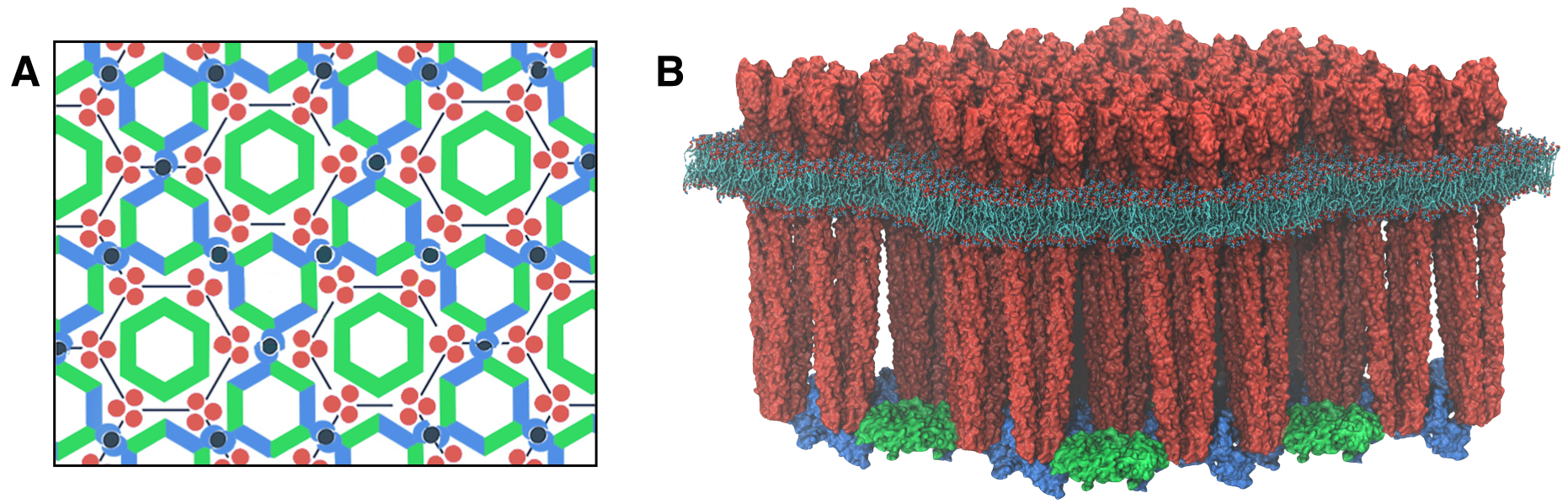
Emergent Features. Central to their chemotactic ability, chemoreceptors cluster along with CheA and CheW to form highly-ordered, supramolecular complexes known as chemosensory arrays (Figure 2). The extensive molecular interaction networks formed within the chemosensory array place the control of each kinase under the joint regulation of many receptors, giving rise to a highly cooperative signaling response. The cooperativity arising from protein clustering within the array results in a number of exceptional information processing features, including drastic signal amplification and precise adaptation. In particular, bacterial cells can amplify signals more than 50-fold; that is to say, a 2% change in receptor occupancy can bring about a 100% change in the output of the system at the flagellar motors. This feature allows cells to sense minute changes in concentration—less than three molecules per cell volume! In addition, precise adaption (i.e., the ability of the chemosensory system to return exactly to its pre-stimulus activity), allows the cell to respond sensitively to chemical gradients over an incredibly wide range of background chemical concentrations—up to five orders of magnitude in size. Such high-performance signaling features are reminiscent of those found in the signaling assemblies of more complex eukaryotic cells such as neurons and lymphocytes. Indeed, a loose analogy can be drawn between the mechanically-based sensational apparatus of a bacterium and the electrochemically-based machinery of higher organisms: the chemosensory array serves as the sensory organ and brain of the organism while the downstream network as a whole provides a type of centralized nervous system, which is used to bias the organism's motor output based on sensory data.
A fundamental and outstanding problem in the bacterial chemotaxis field concerns an understanding of the particular structural and dynamical characteristics of the chemosensory system giving rise to the aforementioned enhanced signaling features. The past several years have witnessed great strides in the understanding of how the chemosensory array's constituent proteins fit and work together. In particular, trimers of chemoreceptors have been shown to form a universally conserved hexagonal lattice, surrounding interlocked rings of CheA/CheW and CheW-only rings (Figure 2A). Nevertheless, a high-resolution description of the kind needed to explore in detail the molecular mechanisms underlying sensory signal transduction within the array has remained elusive. Recent work at the TCBG has begun to shed light in this area (Figure 2B).
The Chemosensory Array: Structure and Dynamics
A mechanistic understanding of the molecular events underlying signaling within the chemosensory array has been hampered by the lack of a high-resolution characterization of its structure.
Recently, we reported a novel reconstitution of the array, involving the receptor signaling domain, histidine kinase CheA, and adaptor protein CheW. This in vitro system produces thin, nearly crystalline specimens (movie) ideal for high resolution structural analysis of chemosensory arrays by cryo-electron tomography.
Sub-tomogram classification and averaging were subsequently used to produce a density map of the core-signaling unit at 11.3 Angstrom resolution (movie).
Extracting key structural constraints from our density map, computationally techniques such as MDFF were used to construct and refine an atomic model of the core array structure from the thermophillic bacterium Thermotoga maritima, exposing novel interfaces between the component proteins (below, left). In particular, the reported model provides improved knowledge of the positioning of the P3 and P4 domains, incorporates the presence of the CheW only ring, and identifies probable novel side-chain contacts within the extended chemosensory architecture.
Using all-atom molecular dynamics simulations, a distinctive conformational change in CheA was revealed that was interpreted as a key signaling event for chemotaxis in live cells (below, right).
Further work will seek to develop an atomic homology model of the E. coli chemosensory array, including an expansion of the model to include the chemoreceptor transmembrane and periplasmic domains.
CheY Activation
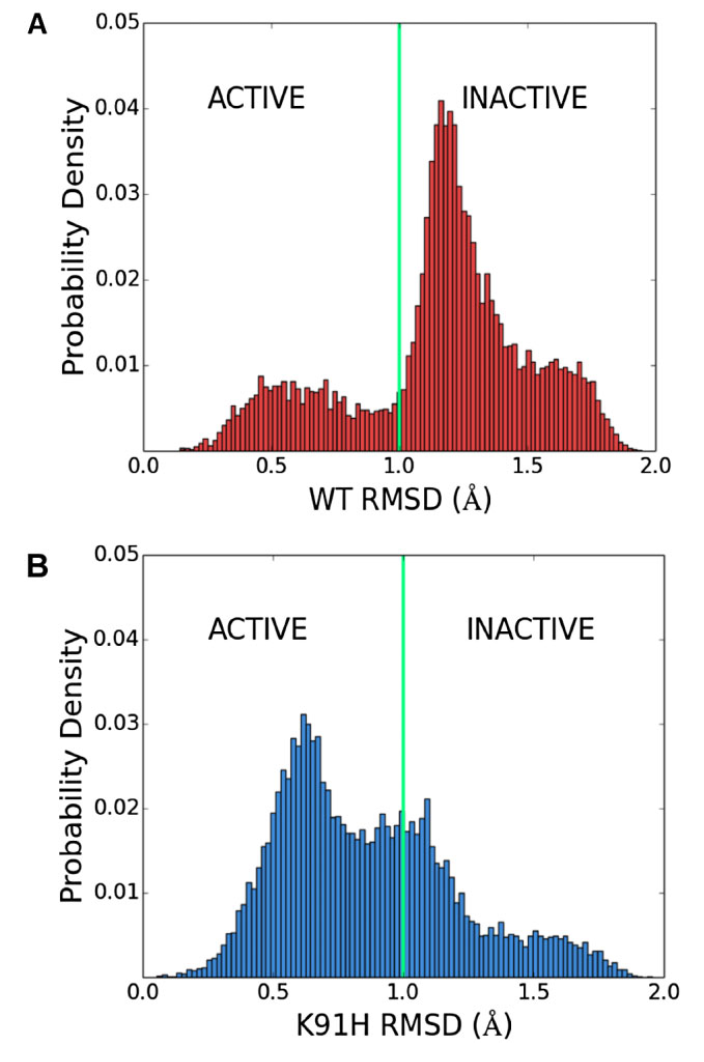
Figure 4: K91 represses the active state of CheY's β4α4 loop. Distribution of RMSD between loop conformations sampled in (A) Wild-type and (B) K91H mutant CheY simulations and an activated β4α4 loop structure (PDB 1FQW) highlights the population shift incurred by the K91H mutation.
Stimulation of E. coli with acetate elevates the acetylation level of the chemotaxis response regulator CheY, enabling it, via an unknown mechanism, to generate clockwise rotation.
We identified K91 and K109 as the main sites of acetylation in vivo and the major source of acetate-dependent clockwise rotation generation.
In addition, we showed that the ability of CheY to generate clockwise rotation was further enhanced by the K91H mutation.
Using molecular dynamics simulations, we show that mutation of K91 drastically affects the conformational dynamics of the β4α4 loop, shifting the loop towards it's activated conformation (Figure 4).
Removal of β4α4 loop repression may represent a general activation mechanism in CheY, pertaining also to the canonical phosphorylation activation pathway.
Publications
- CryoEM and computer simulations reveal a novel kinase conformational switch in bacterial chemotaxis signaling
- CheY's acetylation sites responsible for generating clockwise flagellar rotation in Escherichia coli
*Cassidy C.K., *Himes B.A., *Alvarez J.A., Ma J., Zhao G., Perilla J.R., Schulten K., Zhang P., eLife, 2015
Fraiberg M., Afanzar O., Cassidy C.K., Gabashvili A., Schulten K., Levin Y., Eisenbach M., Molecular Microbiology, 95:231-244, 2015
Additional Resources
Interactive
Interact with the with the chemosensory array's molecular structure and electron density by clicking below.EMD-6319 by Sketchfab For Science on Sketchfab
PDB & EMDB Entries
- PDB: 3JA6 (Core-signaling unit structure from T. maritima)
- EMD: 6319 (CheA2-trimer density map from E. coli)
- EMD: 6320 (CheA2-hexamer density map from E. coli)
News
Investigators
- C. Keith Cassidy - Department of Physics, UIUC, USA
- Klaus Schulten - Department of Physics, UIUC, USA
- Peijun Zhang - Department of Structural Biology, University of Pittsburgh, USA
- Michael Eisenbach - Department of Biological Chemistry, Weizmann Institute of Science, Israel