How to run IMD
How to run Interactive Molecular Dynamics
Step-by-step instructions
-
Obtain a copy of VMD ,
version 1.4b1 or higher, and a copy of
NAMD2 , version 2.1b1
or higher.
-
Append the following lines to your NAMD2 startup file:
IMDon yes IMDport 2030 IMDfreq 1
Assuming your system starts up ok, NAMD will wait for VMD to connect before proceeding with the simulation. IMDport tells NAMD which port to listen on for VMD's connection. IMDfreq tells NAMD to pass coordinates to VMD at every timestep.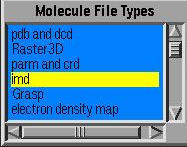
-
Start VMD and load the same system you're simulating in NAMD.
Important: you must load the molecule as an imd molecule. This can be done from the Molecule Files form, selecting imd as the molecule type, or from the command line by typingmol load imd file.psf pdb file.pdb
At present it's necessary to load both a pdb and psf file into VMD, but since these files are also currently required for NAMD this should not pose any new restrictions. -
Assume you are running NAMD on titan , and the IMDport in the
NAMD startup file is 2030. Connect to the simulation by typing the following
from the command line:
imd connect titan 2030
If you are running NAMD on multiple nodes (such as on a Beowulf cluster), you must enter the machine corresponding to node 0 in the simulation. This can be determined by looking at the second line of the output file from NAMD.Other commands to control the connection are:
- imd detach : Detaches from the simulation, leaving it to run.
- imd kill : Detaches from the simulation and tries to terminate it.
- imd keep rate : Sets the rate at which VMD saves simulation frames to the given rate; these frames can be played back at any time using the Animate form, or saved to a DCD file.
- imd transfer rate : Sets the rate at which NAMD sends coordinates to VMD.
- If the simulation crashes or otherwise needs to be restarted, try simply restarting NAMD the same way it was started before. If you get the message Unable to connect to given IMDport , change the IMDport setting in the configuration file to a new value, then try again. (The port will have to be an integer greater than 1024.) You may also have to detach VMD before you can reconnect to the simulation. If you have saved frames from previous simulations, these will be preserved.
Tk widget for IMD
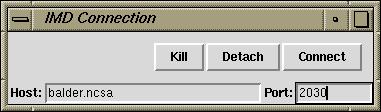 This Tk widget may make more convenient to
connect and disconnect to remote systems. Download the script and
source it from within VMD. Pressing enter after from within
any of the text fields will cause VMD to try to connect to the simulation.
This Tk widget may make more convenient to
connect and disconnect to remote systems. Download the script and
source it from within VMD. Pressing enter after from within
any of the text fields will cause VMD to try to connect to the simulation.
Example scripts
Below are files you can download to test out IMD on your system.alanin.pdb The pdb file.
alanin.psf The psf file.
alanin.conf The NAMD startup file.
alanin.params The Charmm parameter file used by NAMD.
Download all these files into a single directory. Start NAMD using the alanin.conf file, then load an imd molecule using the alanin.psf and alanin.pdb files. Follow the above instructions to connect to the simulation, and watch and watch the action! When you're ready for more fun, you can apply forces using the mouse, or with a 3D tracker.



