Conformational Change of Holliday Junction
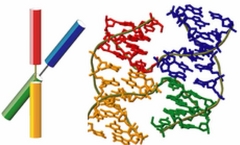
Homologous genetic recombination is essential in maintaining genomic stability such as to protect genomes from double-strand breaks or interstrand crosslinks . It is also important in generating genetic diversity that is necessary for evolution. Holliday junction, a four-way DNA junction, is generally accepted to be the central intermediate in homologous recombination, formed as a result of a reciprocal exchange of DNA strands between two nearly identical DNA molecules. It is named after Robin Holliday who proposed a model of the recombination in 1964.
The Holliday junction is not a static structure. It can undergo spontaneous branch migration in which the branch point can hop forward or backward in a stochastic way; extensive branch migration requires branch migration enzymes to provide a direction over long stretches of DNA. In between of adjacent branch migration events, the Holliday junction can spontaneously fold or unfold, transforming from one stacked conformer to the other. The Holliday junction can be resolved into two duplex DNA molecules via the action of junction resolving enzymes and the extent of genetic information exchange depends on the orientation of resolution.
Conformer Transition Observed from Experiments
The Holliday junction free in solution is structurally polymorphic. Under low salt conditions and in the absence of multivalent ions, the junction extends to an open form, minimizing the repulsion between the negatively charged phosphates concentrated at the junction. In the presence of multivalent cations or in high concentration of monovalent cations, the junction overcomes the electrostatic repulsion and folds into one of two stacked conformers.
Experiments have shown that the conformers interconvert and their relative population depends on the DNA sequence near the junction. Single-molecule measurements from T. Ha's lab, suggested that the conformer transition and the branch migration share the same intermediate, the unfolded open form. Although in physiological condition, the junction folds into the stacked form, upon binding to branch migration enzymes or junction resolving enzymes, it unfolds to various degrees. Therefore, unfolded open forms of the junction appear to be involved in almost every aspect of junction processing. In spite of their potential physiological significance, however, the precise nature of the open form intermediate structure is presently unknown. In physiological conditions, the intermediate forms are too short-lived to be detected directly even in single-molecule measurements, not to mention the knowledge of detailed pathway of conformer transition.
Molecular Dynamics Studies of Conformer Transition
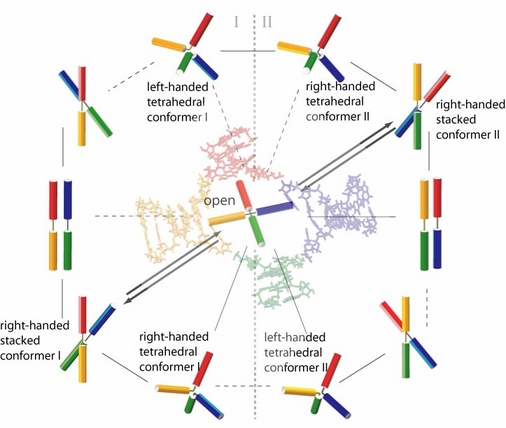
Described in our recent publication, we explore the global conformations involved in the conformer transitions of the Holliday junction mainly utilizing molecular dynamics (MD) simulations (with program NAMD). Facing again with the timescale problem, but in the opposite way as that in experiments, the approachable nanoseconds timescale in current MD is far too short compared with the hundreds of microseconds/milliseconds timescale on which conformer transition happens. Therefore, simulations at an elevated temperature (400 K) were performed to investigate possible transition steps within a schematic framework. The schematic framework was constructed based on available solution structure data, crystal structures, and the principle symmetry of the conformations; a simplified model was introduced in anticipating to catch the most essential global conformational changes of the Holliday junction under conformer transitions.
To be cautious with intermediate forms and pathways deduced from the high temperature simulations, we performed a series of room temperature simulations and used the measured data to explore on a free energy surface governing the Holliday junction solvated in solution. The calculation located various local energy minima, which were actually present in the high temperature simulations. The most probable pathways deduced from MD simulations performed at both elevated and room temperature are practically consistent. In addition to the expected open form intermediates, we observed tetrahedral intermediates which may intrinsically facilitate conformer transition from a purely geometric point of view.
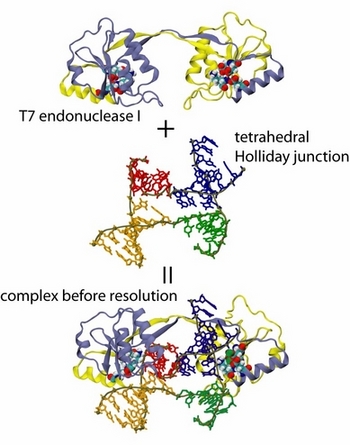
T7 endonuclease I, a junction-resolving enzyme, binds to the Holliday junction with high specificity and makes two cuts diagonally across the junction. In comparative gel electrophoresis experiments, the enzyme-bound junction was seen to adopt two alternative conformations, which correspond well to the right and left -handed tetrahedral intermediates observed in our simulations. A test simulation showed that the tetrahedral form can engage in a stable complex with the enzyme. We note that another endonuclease, E.coli RusA, has also been proposed to bind the Holliday junction in a tetrahedral form. There then arises the interesting possibility that the enzyme recognizes the tetrahedral form populated in a free junction and stabilizes it via binding. In this case the relative populations of the right and left -handed tetrahedral forms may determine the orientation of junction resolution and, hence, the outcome of recombination.
Interesting links of Holliday Junction
- Watch NOVA online "life's greatest miracle" (see the meiosis)
- Schematic strand exchange model, plus a 3D tetrahedral cartoon
- Structure and Folding of Branched Nucleic Acids
- Holliday junction and its related protein complexes
- Current application: DNA branches into nanotech
- Current application: Single molecule nanometronome
Investigators
Page created and maintained by Jin Yu.



