- July 2003 Summer School, Urbana
- June 2004 Workshop in Perth, Australia
- November 2004 Hands-On Workshop in Urbana
- December 2004 Hands-On Workshop in Boston
- May 2005 Hands-On Workshop in Lake Tahoe
- June 2005 Hands-On Workshop in Chicago
- June 2005 Hands-On Workshop in San Francisco
Theoretical and Computational Biophysics Workshops:
Training for the New Discipline

While the life sciences are perhaps the oldest area of human inquiry, the field is still evolving. Today quantitative and computational methods are joining traditional experimental methods, and the combination is opening new areas of research and enabling new discoveries.
"We see the life sciences transformed into a new discipline," says Klaus Schulten, the leader of the Theoretical and Computational Biophysics Group at the University of Illinois at Urbana-Champaign. "We have to take account of this change not just in research but also in education. What is in the research lab today is applied tomorrow in medicine and industry, so we have to train students to deal with quantitative information and computational methods."
Schulten's group, funded as a National Institutes of Health Resource for Macromolecular Modeling and Bioinformatics, is addressing this educational need through hands-on workshops, which have been supported by the National Institutes of Health, the National Science Foundation, and the National Center for Supercomputing Applications. Through these workshops, graduate students, postdocs, and other researchers are able to extend their skill-set to include computational simulation and modeling techniques using the NAMD and VMD software developed by Schulten's group.
Workshops combine theory with hands-on experience
"We really feel that you cannot teach just by lecturing; you have to teach through hands-on examples," explains Schulten. Therefore, the workshops combine lectures on concepts with the practical application of computational methods guided by tutorials that allow participants to explore the examples on their own under the watchful eyes of teaching assistants and lecturers.
"This is the most effective way of learning modeling techniques," says Emad Tajkhorshid, the assistant director of research for the NIH Resource.
After the 2003 workshop, the tutorials, along with all the required data and software, were loaded on laptop computers and provided for participants to use throughout the workshops taught in 2004-2005. This uniform computing environment allows the instructors and workshop participants to avoid obstacles caused by heterogeneous equipment.
The laptops furnished to users also provide portability; by shipping the machines, the workshops can be (and have been) conducted at any location. This enables a wide range of scientists to participate in the workshops.
Hands-On Workshop Series
The Theoretical and Computational Biophysics Group at the University of Illinois launched its workshop series in 2003 with a two-week summer school on the University's Urbana campus. The seven workshops that have been held across the country and even overseas have served more than 200 scientists.
The workshops begin every morning with lectures from experienced researchers, including Klaus Schulten, Zan Luthey-Schulten, and Emad Tajkhorshid. These lectures provide participants with a foundation in the theory behind computational simulation and modeling, examples of the research problems these techniques can effectively address, and descriptions of the use of the software developed by Schulten's group, NAMD and VMD. These programs are widely used in the life sciences community, with more than 60,000 registered users.
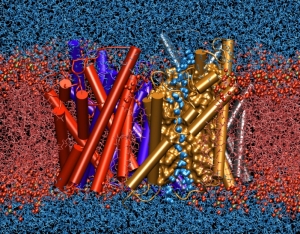
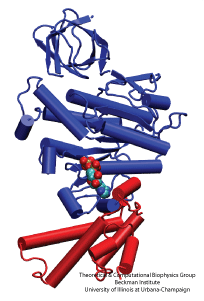
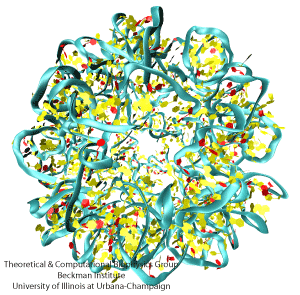
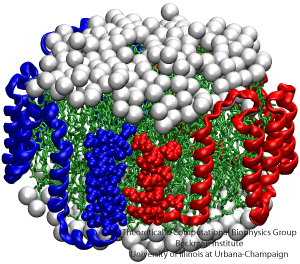
NAMD is a parallel molecular dynamics code designed for high-performance simulation of large biomolecular systems. VMD is a molecular visualization program for displaying, animating, and analyzing large biomolecular systems using 3D graphics. Together, NAMD and VMD form a powerful simulation and visualization environment that is used by tens of thousands of scientists around the world.
"I try to be as practical as possible," says frequent instructor Tajkhorshid, pointing out that the lectures employ numerous examples to connect the topics to real-world research. "I try to walk participants through one concrete project - how we set up a simulation, how we looked at a system, and how we analyzed the results."
Self-directed tutorials provide first-hand experience
The tutorials are the core of the workshop experience. Building on the foundation established through the lectures, the interactive tutorials guide workshop participants through activities that familiarize them with the basics of the NAMD and VMD software and through examples of how to apply the techniques they have learned. The tutorials guide students through tasks that approximate actual research activities.
Because the tutorials are self-directed, students can work at their own pace and focus on topics that most closely correspond to their own research interests.
The first tutorials were developed for the workshop held in the summer of 2003. Graduate students in the group tackled this task because their level of expertise was not so far removed from that of the workshop participants. The experience of learning the software and techniques was fresh in their minds, so they could anticipate the needs and questions of the participants. As a result, the tutorials are accessible and user-friendly.
"The tutorials are based on what we would have wanted to know, what we wish someone had told us," says graduate student and workshop assistant Elizabeth Villa.
Typical lecture schedule
Day 1: Introduction to Protein Structure and Dynamics
- Molecular Graphics Perspective of Protein Structure and Function
- Molecular Dynamics Methods
- Equilibrium Properties of Proteins
- Using NAMD
- Simulated Cooling of Proteins
- Temperature Echoes
- Evolution of Protein Structure
- Sequence and Structure Alignment Algorithms
- Bioinformatics of Aquaporins
- Introduction and Examples
- Introduction to Classical Force Fields
- Methods of Parameterization
- Transport in Aquaporins
- Nanotubes
The tutorials are continually improved based on feedback from workshop participants. "We take great care in asking the participants for their evaluations, using their feedback to refine and improve," Schulten says. "The high quality and the successful execution is due to the tutorials being developed through the workshops over the past few years." New tutorials are regularly added to respond to learners' needs.
The tutorials are posted online, allowing participants to extend their learning beyond the workshops and enabling other learners to participate.
Experienced mentors share their knowledge
Researchers are trailblazers; they tackle novel problems, test novel techniques, and arrive as fresh discoveries. The generous trailblazers led by Schulten then share their knowledge with others.
"I really felt that the learning curve was just too steep," says Luthey-Schulten, a professor of chemistry at the University of Illinois as well as a researcher with the University's Institute for Genomic Biology and the Beckman Institute for Advanced Science and Technology. "Even if you publish papers and tell people how you addressed a problem and got your results, people really need to come and sit next to you and try it themselves with the tools."
And that's just what happens during the workshops. The lectures are given by researchers with expertise in the theory, techniques, and software. The tutorials have been developed and refined by graduate students with first-hand experience conducting computational simulations and visualizing biomolecular systems using NAMD and VMD. And these same graduate students are available during the workshops to answer participants' questions and provide additional support.
Tutorials
All of the tutorials developed by the Theoretical and Computational Biophysics Group at the University of Illinois are available online at http://www.ks.uiuc.edu/Training/Tutorials/.
- Aquaporins with the VMD MultiSeq Tool
- VMD Molecular Graphics
- NAMD Tutorial
- Parameterizing a Novel Residue
- Evolution of Protein Structure Aspartyl-tRNA Synthetase
- Sequence Alignment Algorithms
- Topology File Tutorial
- Stretching Deca-Alanine
- Simulation of Water Permeation through Nanotubes
The graduate students find that the workshops are a learning experience not just for those attending the workshops, but also for those assisting with them. During the workshops, the TAs field novel questions and see problems from new perspectives as they help the participants.
And the questions don't stop when the workshop ends. The partnerships between the mentors and mentees continues as the participants carry their new skills back to their own labs, to be applied to their own research. As questions arise, many will contact their TAs for advice and assistance.
Participants from diverse backgrounds gain new skills
The more than 200 students who have so far participated in the workshops come from diverse backgrounds and have a wide range of research interests. Many are graduate students or post-doctoral researchers, while others have more established careers. Most come from academic and research institutions, but some are from industry. Some are computationally oriented life scientists, while many others are experimentalists who are eager to gain computational skillsto guide their future work. And of course the participants come from across the country and around the world.
"The diversity of the participants really reflects the degree to which the community needs computational tools and modeling," says Tajkhorshid.
Some of the attendees arrive at the workshops with specific research issues with which they need assistance. Perhaps they have been trying to do computational research and have been stymied by technical roadblocks or simply by a lack of time to devote to acquiring the necessary skills. After the workshop, they return to their own labs able to apply the techniques they learned and to transfer them to other researchers.
"We wanted people to be able to start from scratch and end up being able to run a simulation," says Villa, a graduate student who has assisted at several workshops. The goal is to have the participants master the techniques required to set up and run a simulation, and to apply these techniques to their own research interests.
"This is a much faster, more systematic way of bringing students up to speed," Schulten says.
At the end of each day during each workshop, the participants were asked to evaluate the lectures and tutorials. And on the final day of each workshop, the students were asked to complete a more comprehensive evaluation of the workshop as a whole. The workshops have been highly praised by participants, who overall find them relevant, useful, and well-organized.
"When they leave the workshop, they really feel that they've become new kinds of scientists," Schulten says.
Providing clustering skills to enable computational research
The Theoretical and Computational Biophysics Group complements its simulation and modeling workshops with workshops on the basics of cluster computing. After all, once researchers have gained computational skills, they need compute systems on which to conduct computational studies.
These cluster computing workshops familiarize users with the basics of designing, building, and deploying clusters of off-the-shelf PC components. In addition to the discussion of the ABC's of clustering, the workshop also includes the opportunity for participants to building a small PC cluster and to test their own applications.
"We would like to help people avoid the mistakes we made with our first cluster," says Jim Phillips, a senior research programmer in the TCB group and one of the cluster workshop instructors.
To that end, Phillips and co-instructor Tim Skirvin, the administrator of the TCB group's in-house clusters, introduce workshop participants to the potential limiting factors that might influence what type of hardware they should install. Do they have a limited amount of space? If so, dual-core processors might allow them to pack more computational punch into a smaller footprint. Is memory bandwidth crucial to their applications? Maybe an SMP system is the way to go. A research group that is considering adding a cluster to its lab needs to consider how much space is available, how much current the cluster will draw, how the hardware will be cooled, etc.
Cluster Workshops
Cluster workshops have been held on the campus of the University of Illinois at Urbana-Champaign on Sept. 22-23, 2005, and Nov. 10-11, 2005.