João Ribeiro

The focus of my career has been to combine biology/biochemistry with computer science. My research focus is on improving the usability of computational chemistry tools to overcome the initial learning curve barrier and computational technical difficulties (e.g., use of scripting languages), hindering the general use of these programs.
As the leading developer of structure preparation tools in VMD, like QwikMD and Molefacture, my focus is to develop tools that any user can employ on their research, independent on the level of expertise in using computational modeling programs.
Home Department: Beckman Institute for Advanced Science and Technology.
Office Address: Room 3055, Beckman Institute, 405 N. Mathews Avenue, Urbana, Illinois 61801.
Office Phone: (217) 300-8728
Email Address: jribeiro@illinois.edu
LinkedIn page: https://www.linkedin.com/in/joaovieiraribeiro/
Education
- PhD in Chemistry (2014) - - Faculty of Sciences, University of Porto - Portugal
Advisor: Dra. Maria João Ramos
- Diploma Degree in Bioinformatics (2009) - - Biotechnology Superior School, Universidade Católica Portuguesa - Portugal
Advisor: Dra. Célia Manaia
Research Interests
- Computational Modeling
- Drug Design
- Protein-Protein interactions
- Molecular Biology
- Molecular Docking
- Molecular Dynamics
- Molecular Structure Preparation
- Molecular Visualization

QwikMD - Gateway to Easy Simulation
To assist experimentalists and any novice to MD to overcome the initial learning currve barrier of MD simulation software, we developed QwikMD, a user interface that connects the widely employed and user-friendly molecular graphics program VMD to the widely adopted MD program NAMD.
By enabling easy control of every step, QwikMD, meets even the needs of experts in the field, increasing the efficiency and quality of their work.
Read More
QwikMD offers:
Easy Setup of MD Simulations
Point Mutations
Changes in Protonation
Protocols for MD and SMD
Live View Simulations
Integrated Basic Analysis
Info Buttons to Guide Novices
Advanced Protocols
Membrane Environment
Advanced Analysis
Available on Amazon Cloud
Full log Capabilities
Easy Reproducibility
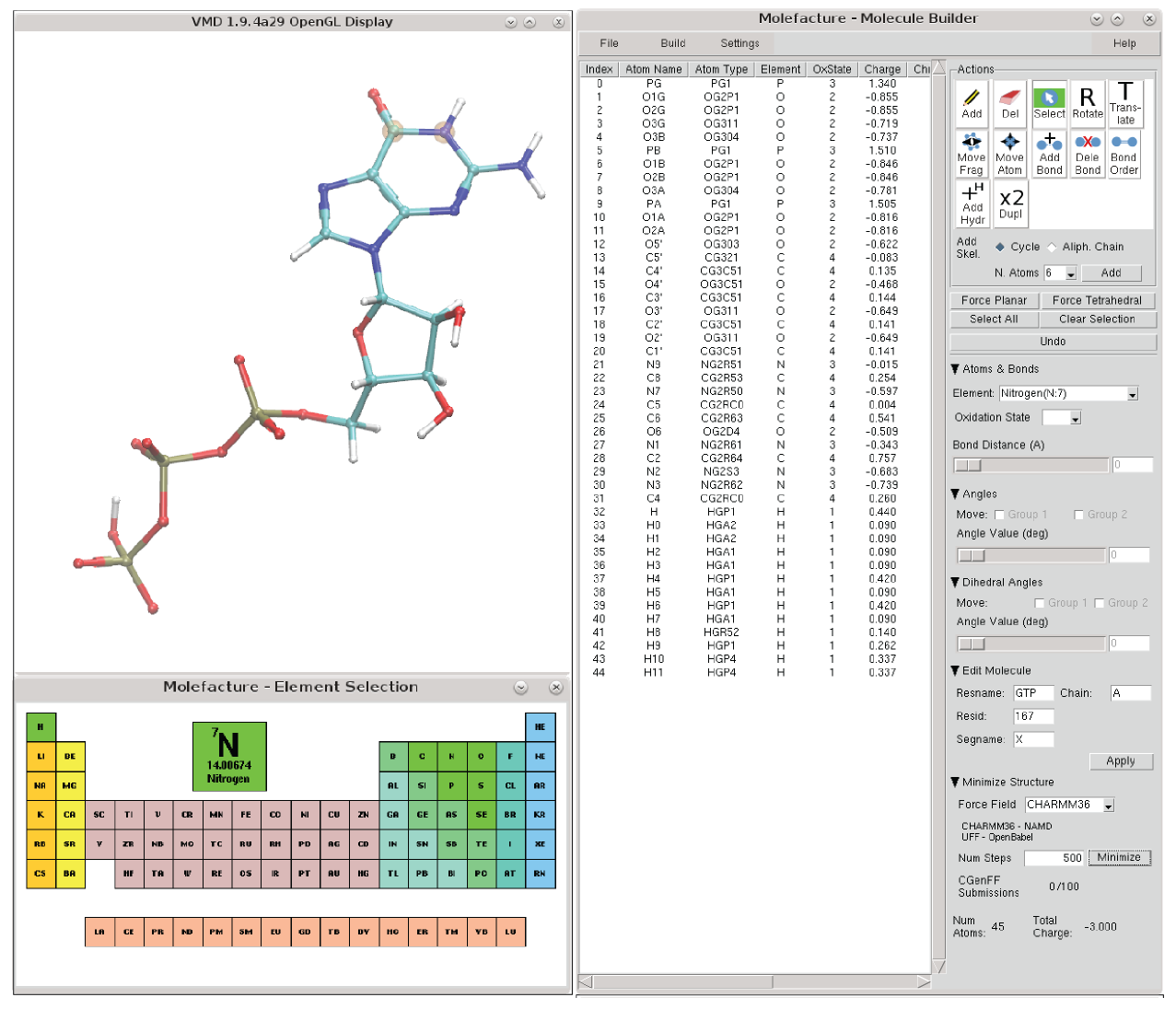
Molefacture - Small Molecule Modeling
The Molefacture plugin provides VMD users with an interface to create and edit molecules. This includes the ability to add, delete, or manipulate their structure at an atomic level, and to build new components from a library of common fragments. Molefacture also interfaces with the CGenFF webserver to retrieve molecular topology and parameters, exporting mol2 file ready to be submitted, and reading in the resulting streaming file (.str).
Read More
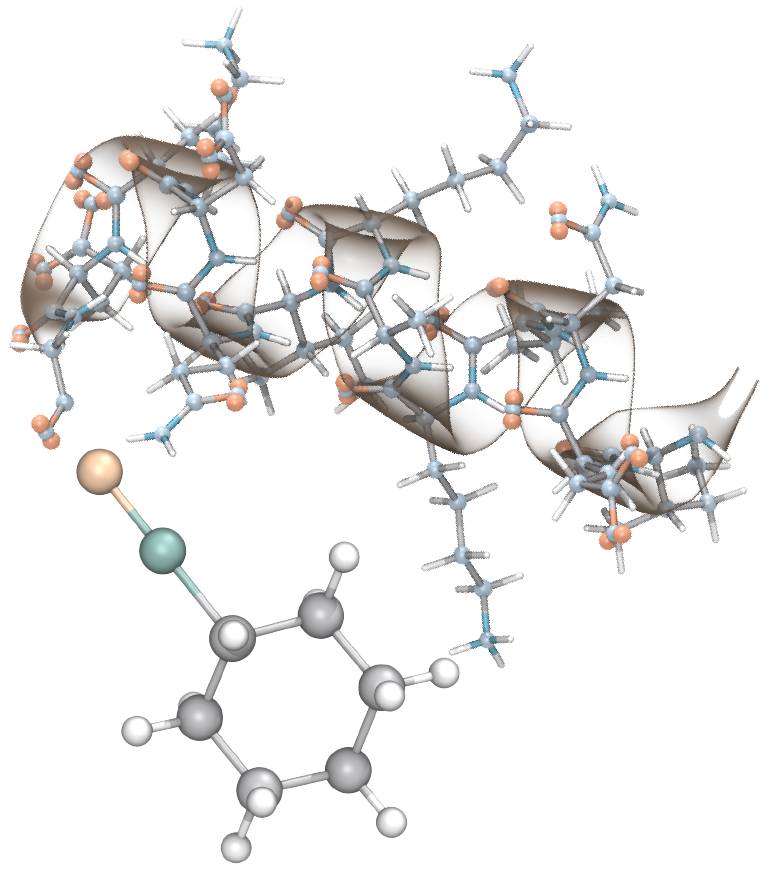
PSFGEN - Simulation Structure Preparation
The version 2.0 of psfgen was extensively modified and improved to meet the current standards in the size of the structures, and the modern versions of additive CHARMM force-field, and polarizable DRUDE force (please see CHARMM Force-Field web page).
Read More
New Functionalities:
Hydrogen Mass Repartition
Structure Preparation for Drude Force Field
Structure Preparation Containing Colinear Lone Pairs
Psfgen Formal log file to store all the information printed to the console
Previous Work
My previous work was dedicated to the development of VMD plugins aiming to aid the user in the use commonly applied computational chemistry techniques, such as Molecular Docking (vsLab), Molecular Volume and Surface calculation(VolArea), Computational Alanine Scanning Mutagenesis(compASM) and to locate chemical motifs in protein structures(Chem-Path-Tracker)
ArtWork
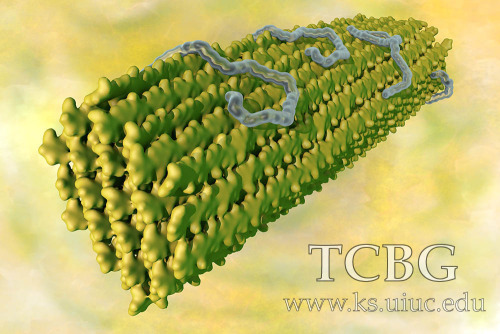


Publications
- NAMD goes quantum: an integrative suite for hybrid simulations; M.C.R.Melo, R.C. Bernardi, T. Rudack, M. Scheurer, C. Riplinger, J.C. Phillips, J.D.C. Maia, G. B. Rocha, J.V. Ribeiro, J.E. Stone, F. Neese, K. Schulten, Z. Luthey-Schulten; Nature Methods, (2018)
- QwikMD-Gateway for Easy Simulation with VMD and NAMD; JV Ribeiro,RC Bernardi, T Rudack, K Schulten, E Tajkhorshid, Biophysical Journal. 2018
- Making Classical and Hybrid (QM/MM) Molecular Dynamics Easy and Fast with QwikMD; JV Ribeiro,RC Bernardi, T Rudack, K Schulten, Biophysical Journal. 2017
- QwikMD Integrative Molecular Dynamics Toolkit for Novices and Experts; JV Ribeiro, RC Bernardi, T Rudack, JE Stone ,JC Phillips,PL Freddolino, K Schulten; Scientific Reports. 2016
- Atomic detail visualization of photosynthetic membranes with GPU-accelerated ray tracing; JE Stone, M Sener, KL Vandivort, A Barragan, A Singharoy, I Teo, JV Ribeiro, B Isralewitz, B Liu, BC Goh, JC Phillips, C MacGregor-Chatwin, MP Johnson, LF Kourkoutis, CN Hunter, K Schulten, Parallel Computing. 2016
- Easy and Fast Setup of Molecular Dynamics Simulations: Combining VMD and NAMD for Experimentalists; JV Ribeiro,RC Bernardi, T Rudack, K Schulten, Biophysical Journal. 2016
- Chem-Path-Tracker: An automated tool to analyze chemical motifs in molecular structures; JV Ribeiro, NMFSA Cerqueira PA Fernandes and MJ Ramos, Chem Biol Drug Des. 2014 (Cover)
- Volarea - A bioinformatics tool to calculate the surface area and the volume of molecular systems;JV Ribeiro, JAC Tamames, NMFSA Cerqueira PA Fernandes and MJ Ramos, Chem Biol Drug Des. 2013
- CompASM: an Amber-VMD alanine scanning mutagenesis plug-in; JV Ribeiro, NMFSA Cerqueira, I Moreina, PA Fernandes and MJ Ramos, Theor Chem Acc 2012
- VsLab-An implementation for virtual high-throughput screening using AutoDock and VMD; NMFSA Cerqueira, J Ribeiro, PA Fernandes and MJ Ramos, Quantum Chem 2011
Presentations
Oral Presentations
Data Standardization in Simulations, MolSSI workshop: Molecular Dynamics Software Interoperability, Brooklyn, NY, USA, 2019.
Accessible Molecular Modeling Environment with VMD and NAMD, ACS National Meeting, San Diego, CA, USA, 2019.
Molecular Dynamics Simulations with NAMD and VMD, PRACE Seasonal School 2019, Sweden - HPC for Life Sciences at Stockholm, Sweden, 2019.
Molecular Dynamics Simulations with NAMD and VMD, "Hands-on" Workshop on Computational Biophysics at Pittsburgh, PA, 2019.
Introduction to Easy and Fast Simulations with QwikMD, "Hands-on" Workshop on Computational Biophysics at Pittsburgh, PA, 2018.
Introduction to QwikMD and Amazon Cloud, "Hands-on" Workshop on Computational Biophysics at Urbana, IL, 2017.
Introduction to QwikMD, "Hands-on" Workshop on Computational Biophysics at Urbana, IL, 2016.
Usage of Poly(NIPAm) in the biofuel production, Publication Number: 292, Polymer Modeling: Structure, Dynamics and Function Section, 249th ACS National Meeting & Exposition, Denver CO, USA, 2015.
Poster Presentations
QwikMD-Gateway for Easy Simulation with VMD and NAMD, 256th ACS National Meeting, Boston MA, USA, 2018.
QwikMD-Gateway for Easy Simulation with VMD and NAMD, 62th Annual Meeting Biophysical Society, San Francisco CA, USA, 2018.
Making Classical and Hybrid (QM/MM) Molecular Dynamics Easy and Fast with QwikMD, 61th Annual Meeting Biophysical Society, New Orleans LA, USA, 2017.
Easy and Fast Molecular Dynamics Simulations for Novices and Experts – QwikMD, 60th Annual Meeting Biophysical Society, Los Angeles CA, USA, 2016.



