#include <colvarcomp.h>
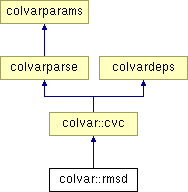
Inheritance diagram for colvar::rmsd:

Public Methods | |||
| rmsd (std::string const &conf) | |||
| Constructor. More... | |||
| virtual | ~rmsd () | ||
| virtual void | calc_value () | ||
| \brief Calculate the variable. More... | |||
| virtual void | calc_gradients () | ||
| \brief Calculate the atomic gradients, to be reused later in order to apply forces. More... | |||
| virtual void | calc_force_invgrads () | ||
| \brief Calculate the total force from the system using the inverse atomic gradients. More... | |||
| virtual void | calc_Jacobian_derivative () | ||
| \brief Calculate the divergence of the inverse atomic gradients. More... | |||
| virtual void | apply_force (colvarvalue const &force) | ||
\brief Apply the collective variable force, by communicating the atomic forces to the simulation program (Note: the ft member is not altered by this function) Note: multiple calls to this function within the same simulation step will add the forces altogether
| |||
| virtual cvm::real | dist2 (colvarvalue const &x1, colvarvalue const &x2) const | ||
| \brief Square distance between x1 and x2 (can be redefined to transparently implement constraints, symmetries and periodicities) colvar::cvc::dist2() and the related functions are declared as "const" functions, but not "static", because additional parameters defining the metrics (e.g. the periodicity) may be specific to each colvar::cvc object. If symmetries or periodicities are present, the colvar::cvc::dist2() should be redefined to return the "closest distance" value and colvar::cvc::dist2_lgrad(), colvar::cvc::dist2_rgrad() to return its gradients. If constraints are present (and not already implemented by any of the colvarvalue types), the colvar::cvc::dist2_lgrad() and colvar::cvc::dist2_rgrad() functions should be redefined to provide a gradient which is compatible with the constraint, i.e. already deprived of its component normal to the constraint hypersurface. Finally, another useful application, if you are performing very many operations with these functions, could be to override the colvarvalue member functions and access directly its member data. For instance: to define dist2(x1,x2) as (x2.real_value-x1.real_value)*(x2.real_value-x1.real_value) in case of a scalar colvarvalue type. More... | |||
| virtual colvarvalue | dist2_lgrad (colvarvalue const &x1, colvarvalue const &x2) const | ||
| \brief Gradient(with respect to x1) of the square distance (can be redefined to transparently implement constraints, symmetries and periodicities). More... | |||
| virtual colvarvalue | dist2_rgrad (colvarvalue const &x1, colvarvalue const &x2) const | ||
| \brief Gradient(with respect to x2) of the square distance (can be redefined to transparently implement constraints, symmetries and periodicities). More... | |||
Protected Attributes | |||
| cvm::atom_group * | atoms | ||
| Atom group. More... | |||
| std::vector< cvm::atom_pos > | ref_pos | ||
| Reference coordinates (for RMSD calculation only) Includes sets with symmetry permutations (n_permutations * n_atoms). More... | |||
| size_t | n_permutations | ||
| Number of permutations of symmetry-related atoms. More... | |||
| size_t | best_perm_index | ||
| Index of the permutation yielding the smallest RMSD (0 for identity). More... | |||
Definition at line 1425 of file colvarcomp.h.
|
|
Constructor.
|
|
|
Definition at line 1446 of file colvarcomp.h. |
|
|
\brief Apply the collective variable force, by communicating the atomic forces to the simulation program (Note: the ft member is not altered by this function) Note: multiple calls to this function within the same simulation step will add the forces altogether
Implements colvar::cvc. Definition at line 1135 of file colvarcomp_distances.C. References atoms, force, and colvarvalue::real_value. |
|
|
\brief Calculate the total force from the system using the inverse atomic gradients.
Reimplemented from colvar::cvc. Definition at line 1142 of file colvarcomp_distances.C. References atoms, colvar::cvc::ft, and colvarvalue::real_value. |
|
|
\brief Calculate the atomic gradients, to be reused later in order to apply forces.
Reimplemented from colvar::cvc. Definition at line 1121 of file colvarcomp_distances.C. References atoms, best_perm_index, colvarmodule::real, colvarvalue::real_value, ref_pos, and colvar::cvc::x. |
|
|
\brief Calculate the divergence of the inverse atomic gradients.
Reimplemented from colvar::cvc. Definition at line 1156 of file colvarcomp_distances.C. References atoms, colvardeps::f_ag_center, colvardeps::f_ag_rotate, colvar::cvc::jd, colvarvalue::real_value, ref_pos, and colvar::cvc::x. |
|
|
\brief Calculate the variable.
Implements colvar::cvc. Definition at line 1094 of file colvarcomp_distances.C. References atoms, best_perm_index, n_permutations, colvarvalue::norm2, colvarmodule::real, colvarvalue::real_value, ref_pos, colvarmodule::sqrt, colvar::cvc::value, and colvar::cvc::x. |
|
||||||||||||
|
\brief Square distance between x1 and x2 (can be redefined to transparently implement constraints, symmetries and periodicities) colvar::cvc::dist2() and the related functions are declared as "const" functions, but not "static", because additional parameters defining the metrics (e.g. the periodicity) may be specific to each colvar::cvc object. If symmetries or periodicities are present, the colvar::cvc::dist2() should be redefined to return the "closest distance" value and colvar::cvc::dist2_lgrad(), colvar::cvc::dist2_rgrad() to return its gradients. If constraints are present (and not already implemented by any of the colvarvalue types), the colvar::cvc::dist2_lgrad() and colvar::cvc::dist2_rgrad() functions should be redefined to provide a gradient which is compatible with the constraint, i.e. already deprived of its component normal to the constraint hypersurface. Finally, another useful application, if you are performing very many operations with these functions, could be to override the colvarvalue member functions and access directly its member data. For instance: to define dist2(x1,x2) as (x2.real_value-x1.real_value)*(x2.real_value-x1.real_value) in case of a scalar colvarvalue type.
Reimplemented from colvar::cvc. |
|
||||||||||||
|
\brief Gradient(with respect to x1) of the square distance (can be redefined to transparently implement constraints, symmetries and periodicities).
Reimplemented from colvar::cvc. |
|
||||||||||||
|
\brief Gradient(with respect to x2) of the square distance (can be redefined to transparently implement constraints, symmetries and periodicities).
Reimplemented from colvar::cvc. |
|
|
Atom group.
Definition at line 1431 of file colvarcomp.h. Referenced by apply_force, calc_force_invgrads, calc_gradients, calc_Jacobian_derivative, and calc_value. |
|
|
Index of the permutation yielding the smallest RMSD (0 for identity).
Definition at line 1441 of file colvarcomp.h. Referenced by calc_gradients, and calc_value. |
|
|
Number of permutations of symmetry-related atoms.
Definition at line 1438 of file colvarcomp.h. Referenced by calc_value. |
|
|
Reference coordinates (for RMSD calculation only) Includes sets with symmetry permutations (n_permutations * n_atoms).
Definition at line 1435 of file colvarcomp.h. Referenced by calc_gradients, calc_Jacobian_derivative, and calc_value. |
 1.2.14 written by Dimitri van Heesch,
© 1997-2002
1.2.14 written by Dimitri van Heesch,
© 1997-2002