



Next: Angles
Up: Defining collective variables
Previous: Choosing a function
Contents
Index
Subsections
The distance {...} block defines a distance component between the two atom groups, group1 and group2.
List of keywords (see also ![[*]](crossref.png) for additional options):
for additional options):
-
group1
 First group of atoms
First group of atoms
Context: distance
Acceptable values: Block group1 {...}
Description: First group of atoms.
-
group2: analogous to group1
-
forceNoPBC
 Calculate absolute rather than minimum-image distance?
Calculate absolute rather than minimum-image distance?
Context: distance
Acceptable values: boolean
Default value: no
Description: By default, in calculations with periodic boundary conditions, the
distance component returns the distance according to the
minimum-image convention. If this parameter is set to yes,
PBC will be ignored and the distance between the coordinates as maintained
internally will be used. This is only useful in a limited number of
special cases, e.g. to describe the distance between remote points
of a single macromolecule, which cannot be split across periodic cell
boundaries, and for which the minimum-image distance might give the
wrong result because of a relatively small periodic cell.
-
oneSiteTotalForce
 Measure total force on group 1 only?
Measure total force on group 1 only?
Context: angle, dipoleAngle, dihedral
Acceptable values: boolean
Default value: no
Description: If this is set to yes, the total force is measured along
a vector field (see equation (13.25) in
section ![[*]](crossref.png) ) that only involves atoms of
group1. This option is only useful for ABF, or custom
biases that compute total forces. See
section
) that only involves atoms of
group1. This option is only useful for ABF, or custom
biases that compute total forces. See
section ![[*]](crossref.png) for details.
for details.
The value returned is a positive number (in Å), ranging from 0
to the largest possible interatomic distance within the chosen
boundary conditions (with PBCs, the minimum image convention is used
unless the forceNoPBC option is set).
The distanceZ {...} block defines a distance projection
component, which can be seen as measuring the distance between two
groups projected onto an axis, or the position of a group along such
an axis. The axis can be defined using either one reference group and
a constant vector, or dynamically based on two reference groups.
One of the groups can be set to a dummy atom to allow the use of an absolute Cartesian coordinate.
List of keywords (see also ![[*]](crossref.png) for additional options):
for additional options):
This component returns a number (in Å) whose range is determined
by the chosen boundary conditions. For instance, if the 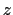 axis is
used in a simulation with periodic boundaries, the returned value ranges
between
axis is
used in a simulation with periodic boundaries, the returned value ranges
between 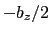 and
and 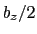 , where
, where 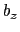 is the box length
along
is the box length
along 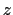 (this behavior is disabled if forceNoPBC is set).
(this behavior is disabled if forceNoPBC is set).
The distanceXY {...} block defines a distance projected on
a plane, and accepts the same keywords as the component distanceZ, i.e.
main, ref, either ref2 or axis,
and oneSiteTotalForce. It returns the norm of the
projection of the distance vector between main and
ref onto the plane orthogonal to the axis. The axis is
defined using the axis parameter or as the vector joining
ref and ref2 (see distanceZ above).
List of keywords (see also ![[*]](crossref.png) for additional options):
for additional options):
-
main: see definition of main (distanceZ component)
-
ref: see definition of ref (distanceZ component)
-
ref2: see definition of ref2 (distanceZ component)
-
axis: see definition of axis (distanceZ component)
-
forceNoPBC: see definition of forceNoPBC (distance component)
-
oneSiteTotalForce: see definition of oneSiteTotalForce (distance component)
The distanceVec {...} block defines
a distance vector component, which accepts the same keywords as
the component distance: group1, group2, and
forceNoPBC. Its value is the 3-vector joining the centers
of mass of group1 and group2.
List of keywords (see also ![[*]](crossref.png) for additional options):
for additional options):
-
group1: see definition of group1 (distance component)
-
group2: analogous to group1
-
forceNoPBC: see definition of forceNoPBC (distance component)
-
oneSiteTotalForce: see definition of oneSiteTotalForce (distance component)
The distanceDir {...} block defines
a distance unit vector component, which accepts the same keywords as
the component distance: group1, group2, and
forceNoPBC. It returns a
3-dimensional unit vector
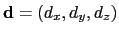 , with
, with
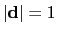 .
.
List of keywords (see also ![[*]](crossref.png) for additional options):
for additional options):
-
group1: see definition of group1 (distance component)
-
group2: analogous to group1
-
forceNoPBC: see definition of forceNoPBC (distance component)
-
oneSiteTotalForce: see definition of oneSiteTotalForce (distance component)
The distanceInv {...} block defines a generalized mean distance between two groups of atoms 1 and 2, weighted with exponent 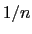 :
:
![$\displaystyle d_{\mathrm{1,2}}^{[n]} \; = \; \left(\frac{1}{N_{\mathrm{1}}N_{\m...
...}\sum_{i,j} \left(\frac{1}{\Vert\mathbf{d}^{ij}\Vert}\right)^{n} \right)^{-1/n}$](img157.png) |
(13.2) |
where
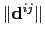 is the distance between atoms
is the distance between atoms 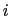 and
and 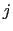 in groups 1 and 2 respectively, and
in groups 1 and 2 respectively, and 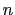 is an even integer.
is an even integer.
List of keywords (see also ![[*]](crossref.png) for additional options):
for additional options):
-
group1: see definition of group1 (distance component)
-
group2: analogous to group1
-
oneSiteTotalForce: see definition of oneSiteTotalForce (distance component)
-
exponent
 Exponent
Exponent 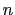 in equation 13.2
in equation 13.2
Context: distanceInv
Acceptable values: positive even integer
Default value: 6
Description: Defines the exponent to which the individual distances are elevated before averaging. The default value of 6 is useful for example to applying restraints based on NOE-measured distances.
This component returns a number in Å, ranging from 0
to the largest possible distance within the chosen boundary conditions.




Next: Angles
Up: Defining collective variables
Previous: Choosing a function
Contents
Index
vmd@ks.uiuc.edu
![[*]](crossref.png) for additional options):
for additional options):
![[*]](crossref.png) for additional options):
for additional options):
![[*]](crossref.png) ) that only involves atoms of
group1. This option is only useful for ABF, or custom
biases that compute total forces. See
section
) that only involves atoms of
group1. This option is only useful for ABF, or custom
biases that compute total forces. See
section ![[*]](crossref.png) for details.
for details.
![[*]](crossref.png) for additional options):
for additional options):
![[*]](crossref.png) for additional options):
for additional options):
![[*]](crossref.png) for additional options):
for additional options):
![]() , with
, with
![]() .
.
![[*]](crossref.png) for additional options):
for additional options):
![]() :
:
![[*]](crossref.png) for additional options):
for additional options):