From: Aashish Bhatt (aashish.ph16221_at_inst.ac.in)
Date: Thu Oct 08 2020 - 08:43:44 CDT
Dear Sir/ma'am
I have one system Alpha-cyclodextrin with peptide. I want to study
host-guest interaction.
for this study, I am following the paper.
https://pubs.acs.org/doi/pdf/10.1021/jp3128324
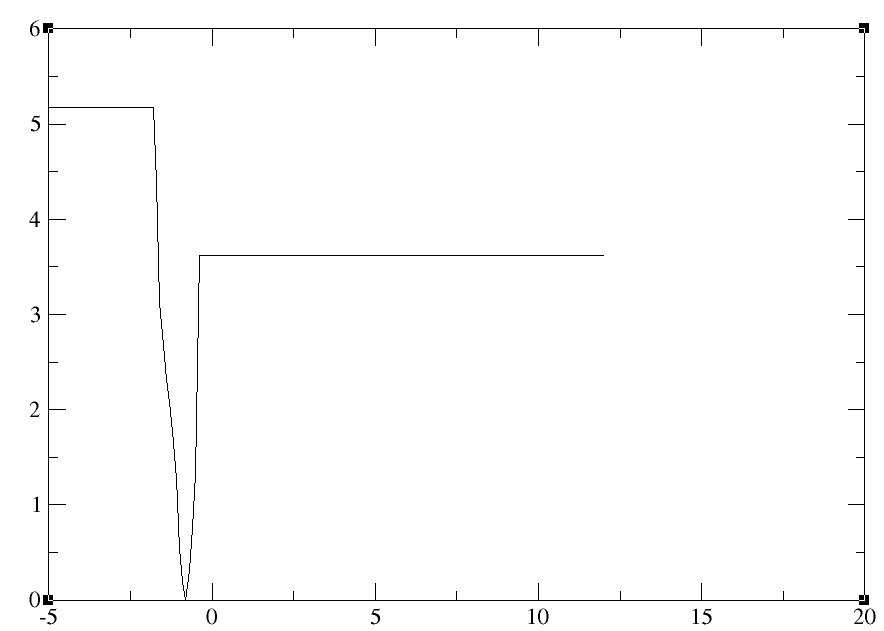
Whenever I performing the Adaptive biasing force method I am getting the
following type of graph for Pmf(Kcal/mol) vs zeta.
[image: image.png]
I have chosen the heavy atoms of the peptide whose inside the
alpha-cyclodextrin as the main atoms and second atoms are oxygen
atoms present in the glycosidic bond.
I have also query ho to choose lowerboundary and upperboundary.
I have also attached the colvar file.
Kindly give me some suggestions to understand this issue.
Best Regards
Aashish

This archive was generated by hypermail 2.1.6 : Thu Dec 31 2020 - 23:17:14 CST