



Next: A crash course
Up: NAMD 2.13b1 User's Guide
Previous: External Program Forces
Contents
Index
Collective Variable-based Calculations (Colvars)1
This chapter can also be downloaded as a separate manual (PDF and HTML) at the webpage:
http://colvars.github.io
In molecular dynamics simulations, it is often useful to reduce the large number of degrees of freedom of a physical system into few parameters whose statistical distributions can be analyzed individually, or used to define biasing potentials to alter the dynamics of the system in a controlled manner.
These have been called `order parameters', `collective variables', `(surrogate) reaction coordinates', and many other terms.
Here we use primarily the term `collective variable' (shortened to colvar), which indicates any differentiable function of atomic Cartesian coordinates,
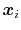 , with
, with 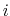 between
between 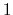 and
and 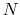 , the total
number of atoms:
, the total
number of atoms:
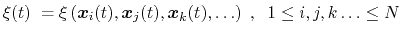 |
(35) |
The Colvars module in NAMD may be used in both MD simulations and energy minimization runs.
The module is designed to perform multiple tasks concurrently during or after a simulation, the most common of which are:
- apply restraints or biasing potentials to multiple colvars, tailored on the system by choosing from a wide set of basis functions, without limitations on their number or on the number of atoms involved; while this can in principle be done through a TclForces script, using the Colvars module is both easier and computationally more efficient;
- calculate potentials of mean force (PMFs) along any set of colvars, using different enhanced sampling methods, such as Adaptive Biasing Force (ABF), metadynamics, steered MD and umbrella sampling; variants of these methods that make use of an ensemble of replicas are supported as well;
- calculate statistical properties of the colvars, such as running averages and standard deviations, correlation functions of pairs of colvars, and multidimensional histograms: this can be done either at run-time without the need to save very large trajectory files, or after a simulation has been completed using VMD and the cv command or NAMD and the coorfile read command as illustrated in 17.
Detailed explanations of the design of the Colvars module are provided in reference [27]. Please cite this reference whenever publishing work that makes use of this module.
Subsections




Next: A crash course
Up: NAMD 2.13b1 User's Guide
Previous: External Program Forces
Contents
Index
http://www.ks.uiuc.edu/Research/namd/
![]() , with
, with ![]() between
between ![]() and
and ![]() , the total
number of atoms:
, the total
number of atoms: