



Next: Selecting atoms for colvars:
Up: Collective Variable-based Calculations (Colvars)1
Previous: General parameters and input/output
Contents
Index
Subsections
Defining collective variables and their properties
In the configuration file each colvar is defined by the keyword
colvar, followed by its configuration options within curly braces: colvar { ... }. One of these options is the name of a colvar component: for example, including rmsd { ... } defines the colvar as a RMSD function. In most applications, only one component is used, and the component is equal to the colvar.
The full list of colvar components can be found in Section 10.4, with the syntax to select atoms in Section 10.3.
The following section lists several options to control the behavior of a single colvar, regardless of its type.
General options for a collective variable
The following options are not required by default; however, the first four are very frequently used:
- name
 Name of this colvar
Name of this colvar 
Context: colvar
Acceptable Values: string
Default Value: ``colvar'' + numeric id
Description: The name is an unique case-sensitive string which allows the
Colvars module to identify this colvar unambiguously; it is also
used in the trajectory file to label to the columns corresponding
to this colvar.
- width
 Colvar fluctuation scale, or resolution for grid-based methods
Colvar fluctuation scale, or resolution for grid-based methods 
Context: colvar
Acceptable Values: positive decimal
Default Value: 1.0
Description: This number has the same physical unit as the colvar value and defines an effective colvar unit.
Biasing algorithms use it for different purposes.
Harmonic restraints (10.5.4) use it to set the physical unit of the force constant, which is useful for multidimensional restraints involving colvars with different units or scale which may then be defined by a single, scaled force constant.
Histogram (10.5.7) and ABF biases (10.5.1) interpret it as the grid spacing in the direction of this variable.
Metadynamics (10.5.3) uses it to set the width of newly added hills.
In other cases, it is simplest to keep the default value of 1, so that harmonic force constants are provided in their usual physical unit.
When a non-unity width is required by the application, the optimal value is application-dependent, but can often be thought of as a user-provided estimate of the fluctuation amplitude for the colvar.
In those cases, it should generally be set smaller than or equal to the standard deviation of the colvar during a very short simulation run.
- lowerBoundary
 Lower boundary of the colvar
Lower boundary of the colvar 
Context: colvar
Acceptable Values: decimal
Description: Defines the lowest end of the interval of ``relevant'' values for the colvar.
This number can be either a true physical boundary, or a user-defined number.
Together with upperBoundary and width, it is used to define a grid of values along the colvar (not available for colvars based on distanceDir, distanceVec, and orientation).
This option does not affect dynamics: to confine a colvar within a certain interval, the options lowerWall and lowerWallConstant should be used.
- upperBoundary
 Upper boundary of the colvar
Upper boundary of the colvar 
Context: colvar
Acceptable Values: decimal
Description: Similarly to lowerBoundary, defines the highest possible or allowed value.
- hardLowerBoundary
 Whether the lower boundary is the physical lower limit
Whether the lower boundary is the physical lower limit 
Context: colvar
Acceptable Values: boolean
Default Value: off
Description: This option does not affect simulation results, but enables some internal optimizations.
Depending on its mathematical definition, a colvar may have ``natural'' boundaries: for example, a distance colvar has a ``natural'' lower boundary at 0. Setting this option instructs the Colvars module that the user-defined lower boundary is ``natural''.
See Section 10.4 for the physical ranges of values of each component.
- hardUpperBoundary
 Whether the upper boundary is the physical upper limit of the colvar's values
Whether the upper boundary is the physical upper limit of the colvar's values 
Context: colvar
Acceptable Values: boolean
Default Value: off
Description: Analogous to hardLowerBoundary.
- expandBoundaries
 Allow to expand the two boundaries if needed
Allow to expand the two boundaries if needed 
Context: colvar
Acceptable Values: boolean
Default Value: off
Description: If defined, biasing and analysis methods may keep their own copies
of lowerBoundary and upperBoundary, and expand
them to accommodate values that do not fit in the initial range.
Currently, this option is used by the metadynamics bias
(10.5.3) to keep all of its hills fully within
the grid. This option cannot be used when
the initial boundaries already span the full period of a periodic
colvar.
- subtractAppliedForce
 Do not include biasing forces in the total force for this colvar
Do not include biasing forces in the total force for this colvar 
Context: colvar
Acceptable Values: boolean
Default Value: off
Description: If the colvar supports total force calculation (see 10.4.3), all forces applied to this colvar by biases will be removed from the total force.
This keyword allows to recover some of the ``system force'' calculation available in the Colvars module before version 2016-08-10.
Please note that removal of all other external forces (including biasing forces applied to a different colvar) is no longer supported, due to changes in the underlying simulation engines (primarily NAMD).
This option may be useful when continuing a previous simulation where the removal of external/applied forces is essential.
For all new simulations, the use of this option is not recommended.
The following options are useful to define restraints (confining potentials) for this colvar.
To apply moving restraints, or restraints to more than one colvar simultaneously, a more convenient option is to use the harmonic bias (10.5.4).
Extended Lagrangian.
The following options enable extended-system
dynamics, where a colvar is coupled to an additional degree of freedom
(fictitious particle) by a harmonic spring.
All biasing and confining forces are then applied to the extended degree
of freedom. The ``actual'' geometric colvar (function of Cartesian
coordinates) only feels the force from the harmonic spring.
This is particularly useful when combined with an ABF bias (10.5.1)
to perform eABF simulations (10.5.2).
- extendedLagrangian
 Add extended degree of freedom
Add extended degree of freedom 
Context: colvar
Acceptable Values: boolean
Default Value: off
Description: Adds a fictitious particle to be coupled to the colvar by a harmonic
spring. The fictitious mass and the force constant of the coupling
potential are derived from the parameters extendedTimeConstant
and extendedFluctuation, described below. Biasing forces on the
colvar are applied to this fictitious particle, rather than to the
atoms directly. This implements the extended Lagrangian formalism
used in some metadynamics simulations [39].
The energy associated with the extended degree of freedom is reported
under the MISC title in NAMD's energy output.
- extendedFluctuation
 Standard deviation between the colvar and the fictitious
particle (colvar unit)
Standard deviation between the colvar and the fictitious
particle (colvar unit) 
Context: colvar
Acceptable Values: positive decimal
Description: Defines the spring stiffness for the extendedLagrangian
mode, by setting the typical deviation between the colvar and the extended
degree of freedom due to thermal fluctuation.
The spring force constant is calculated internally as
 ,
where
,
where 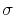 is the value of extendedFluctuation.
is the value of extendedFluctuation.
- extendedTimeConstant
 Oscillation period of the fictitious particle (fs)
Oscillation period of the fictitious particle (fs) 
Context: colvar
Acceptable Values: positive decimal
Default Value: 200
Description: Defines the inertial mass of the fictitious particle, by setting the
oscillation period of the harmonic oscillator formed by the fictitious
particle and the spring. The period
should be much larger than the MD time step to ensure accurate integration
of the extended particle's equation of motion.
The fictitious mass is calculated internally as
 ,
where
,
where 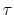 is the period and
is the period and 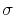 is the typical fluctuation (see above).
is the typical fluctuation (see above).
- extendedTemp
 Temperature for the extended degree of freedom (K)
Temperature for the extended degree of freedom (K) 
Context: colvar
Acceptable Values: positive decimal
Default Value: thermostat temperature
Description: Temperature used for calculating the coupling force constant of the
extended variable (see extendedFluctuation) and, if needed, as a
target temperature for extended Langevin dynamics (see
extendedLangevinDamping). This should normally be left at its
default value.
- extendedLangevinDamping
 Damping factor for extended Langevin dynamics
(ps
Damping factor for extended Langevin dynamics
(ps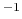 )
) 
Context: colvar
Acceptable Values: positive decimal
Default Value: 1.0
Description: If this is non-zero, the extended degree of freedom undergoes Langevin dynamics
at temperature extendedTemp. The friction force is minus
extendedLangevinDamping times the velocity. This is useful because
the extended dynamics coordinate may heat up in the transient
non-equilibrium regime of ABF. Use moderate damping values, to limit
viscous friction (potentially slowing down diffusive sampling) and stochastic
noise (increasing the variance of statistical measurements). In
doubt, use the default value.
Statistical analysis of collective variables
When the global keyword analysis is defined in the
configuration file, run-time calculations of statistical properties for
individual colvars can be performed. At the moment, several types of
time correlation functions, running averages and running standard
deviations are available.
- corrFunc
 Calculate a time correlation function?
Calculate a time correlation function? 
Context: colvar
Acceptable Values: boolean
Default Value: off
Description: Whether or not a time correlaction function should be calculated
for this colvar.
- corrFuncWithColvar
 Colvar name for the correlation function
Colvar name for the correlation function 
Context: colvar
Acceptable Values: string
Description: By default, the auto-correlation function (ACF) of this colvar,
 , is calculated. When this option is specified, the
correlation function is calculated instead with another colvar,
, is calculated. When this option is specified, the
correlation function is calculated instead with another colvar,
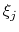 , which must be of the same type (scalar, vector, or
quaternion) as
, which must be of the same type (scalar, vector, or
quaternion) as  .
.
- corrFuncType
 Type of the correlation function
Type of the correlation function 
Context: colvar
Acceptable Values: velocity, coordinate or
coordinate_p2
Default Value: velocity
Description: With coordinate or velocity, the correlation
function
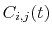 =
=
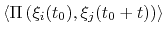 is calculated between
the variables
is calculated between
the variables  and
and 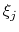 , or their velocities.
, or their velocities.
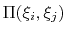 is the scalar product when calculated
between scalar or vector values, whereas for quaternions it is the
cosine between the two corresponding rotation axes. With
coordinate_p2, the second order Legendre polynomial,
is the scalar product when calculated
between scalar or vector values, whereas for quaternions it is the
cosine between the two corresponding rotation axes. With
coordinate_p2, the second order Legendre polynomial,
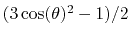 , is used instead of the cosine.
, is used instead of the cosine.
- corrFuncNormalize
 Normalize the time correlation function?
Normalize the time correlation function? 
Context: colvar
Acceptable Values: boolean
Default Value: on
Description: If enabled, the value of the correlation function at 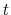 = 0
is normalized to 1; otherwise, it equals to
= 0
is normalized to 1; otherwise, it equals to
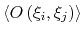 .
.
- corrFuncLength
 Length of the time correlation function
Length of the time correlation function 
Context: colvar
Acceptable Values: positive integer
Default Value: 1000
Description: Length (in number of points) of the time correlation function.
- corrFuncStride
 Stride of the time correlation function
Stride of the time correlation function 
Context: colvar
Acceptable Values: positive integer
Default Value: 1
Description: Number of steps between two values of the time correlation function.
- corrFuncOffset
 Offset of the time correlation function
Offset of the time correlation function 
Context: colvar
Acceptable Values: positive integer
Default Value: 0
Description: The starting time (in number of steps) of the time correlation
function (default: 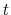 = 0). Note: the value at
= 0). Note: the value at 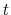 = 0 is always
used for the normalization.
= 0 is always
used for the normalization.
- corrFuncOutputFile
 Output file for the time correlation function
Output file for the time correlation function 
Context: colvar
Acceptable Values: UNIX filename
Default Value:  name
name .corrfunc.dat
.corrfunc.dat
Description: The time correlation function is saved in this file.
- runAve
 Calculate the running average and standard deviation
Calculate the running average and standard deviation 
Context: colvar
Acceptable Values: boolean
Default Value: off
Description: Whether or not the running average and standard deviation should
be calculated for this colvar.
- runAveLength
 Length of the running average window
Length of the running average window 
Context: colvar
Acceptable Values: positive integer
Default Value: 1000
Description: Length (in number of points) of the running average window.
- runAveStride
 Stride of the running average window values
Stride of the running average window values 
Context: colvar
Acceptable Values: positive integer
Default Value: 1
Description: Number of steps between two values within the running average window.
- runAveOutputFile
 Output file for the running average and standard deviation
Output file for the running average and standard deviation 
Context: colvar
Acceptable Values: UNIX filename
Default Value:  name
name .runave.dat
.runave.dat
Description: The running average and standard deviation are saved in this file.




Next: Selecting atoms for colvars:
Up: Collective Variable-based Calculations (Colvars)1
Previous: General parameters and input/output
Contents
Index
http://www.ks.uiuc.edu/Research/namd/