



Next: MARTINI Residue-Based Coarse-Grain Forcefield
Up: Force Field Parameters
Previous: Water Models
Contents
Index
Subsections
Drude polarizable force field
The Drude oscillator model represents induced electronic polarization
by introducing an auxiliary particle attached to each polarizable
atom via a harmonic spring. The advantage with the Drude model is
that it preserves the simple particle-particle Coulomb electrostatic
interaction employed in nonpolarizable force fields,
thus its implementation in NAMD is more straightforward than
alternative models for polarization.
NAMD performs the integration of Drude oscillators
by employing a novel dual Langevin thermostat to freeze the Drude
oscillators while maintaining the warm degrees of freedom
at the desired temperature [37].
Use of the Langevin thermostat enables better parallel scalability
than the earlier reported implementation which made use of
a dual Nosé-Hoover thermostat acting on, and within,
each nucleus-Drude pair [44].
Performance results
show that the NAMD implementation of the Drude model maintains good
parallel scalability,
with an increase in computational cost by not more than twice
that of using a nonpolarizable force field [37].
The Drude polarizable force field requires some extensions to the
CHARMM force field.
The Drude oscillators differ from typical spring bonds only in that they
have an equilibrium length of zero.
The Drude oscillators are optionally supplemented by a maximal
bond length parameter, beyond which a quartic restraining potential
is also applied.
The force field is also extended by an anisotropic spring term
that accounts for out-of-plane forces from a polarized atom and its
covalently bonded neighbor with two more covalently bonded
neighbors (similar in structure to an improper bond).
The screened Coulomb correction of Thole is calculated between
pairs of Drude oscillators that would otherwise be excluded from
nonbonded interaction and
optionally between non-excluded, nonbonded pairs of Drude oscillators
that are within a prescribed cutoff distance [70,71].
Also included in the Drude force field
are the use of off-centered massless interaction sites,
so called ``lone pairs'' (LPs),
to avoid the limitations
of centrosymmetric-based Coulomb interactions [30].
The coordinate of each LP site is constructed based on three ``guide'' atoms.
The calculated forces on the massless LP must be transferred
to the guide atoms,
preserving total force and torque.
After an integration step of velocities and positions,
the position of the LP is updated based on the three guide atoms,
along with additional geometry parameters that give displacement
and in-plane and out-of-plane angles.
See our research web page
(http://www.ks.uiuc.edu/Research/Drude/)
for additional details and parallel performance results.
No additional files are required by NAMD to use the
Drude polarizable force field.
However, it is presently beyond the capability of
the Psfgen tool to generate the PSF file needed to perform
a simulation using the Drude model.
For now, CHARMM is needed to generate correct input files.
The CHARMM force field parameter files
specific to the Drude model are required.
The PDB file must also include
the Drude particles (mass between 0.1 and 1.0)
and the LPs (mass 0).
The Drude particles always immediately follow their parent atom.
The PSF file augments the ``atom'' section with
additional columns that include the
``Thole'' and ``alpha'' parameters for the screened Coulomb
interactions of Thole.
The PSF file also requires additional sections that
list the LPs, including their guide atoms and geometry parameters,
and list the anisotropic interaction terms, including their parameters.
A Drude-compatible PSF file is denoted by the
keyword ``DRUDE'' given along the top line.
The NAMD logging to standard output is extended to provide additional
temperature data on the cold and warm degrees of freedom.
Four additional quantities are listed
on the ETITLE and ENERGY lines:
- DRUDECOM
- gives the temperature for the warm center-of-mass
degrees of freedom,
- DRUDEBOND
- gives the temperature for the cold Drude oscillator
degrees of freedom,
- DRCOMAVG
- gives the average temperature
(averaged since the previously reported temperature)
for the warm center-of-mass degrees of freedom,
- DRBONDAVG
- gives the average temperature
(averaged since the previously reported temperature)
for the cold Drude oscillator degrees of freedom.
The energies resulting from the Drude oscillators and the
anisotropic interactions are summed into the BOND energy.
The energies resulting from the LPs and the screened Coulomb
interactions of Thole are summed into the ELECT energy.
The Drude model should be used with the Langevin thermostat enabled
(Langevin=on).
Doing so permits the use of normal sized time steps (e.g., 1 fs).
The Drude model is also compatible with constant pressure simulation
using the Langevin piston.
Long-range electrostatics may be calculated using PME.
The nonbonded exclusions should generally be set to use either the
1-3 exclusion policy (exclude=1-3)
or the scaled 1-4 exclusion policy (exclude=scaled1-4).
The Drude water model (SWM4-NDP) is a 5-site model
with four charge sites and
a negatively charged Drude particle [43],
with the particles ordered in the input files as
oxygen, Drude particle, LP, hydrogen, hydrogen.
The atoms in the water molecules should be
constrained (rigidBonds=water),
with use of the SETTLE algorithm recommended (useSettle=on).
Explicitly setting the water model (waterModel=swm4) is optional.
- drude
 Perform integration of Drude oscillators?
Perform integration of Drude oscillators? 
Acceptable Values: on or off
Default Value: off
Description: The integration uses a dual Langevin thermostat to freeze the Drude
oscillators while maintaining the warm degrees of freedom at the
desired temperature. Must also enable the Langevin thermostat.
If drude is set to on, then drudeTemp must also be defined.
- drudeTemp
 temperature for freezing the Drude oscillators (K)
temperature for freezing the Drude oscillators (K) 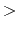
Acceptable Values: non-negative decimal
Description: For stability, the Drude oscillators must be kept at a very cold termpature.
Using a Langevin thermostat, it is possible to set this temperature to 0 K.
- drudeDamping
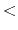 damping coefficient for Drude oscillators (1/ps)
damping coefficient for Drude oscillators (1/ps) 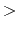
Acceptable Values: positive decimal
Description: The Langevin coupling coefficient to be applied to the Drude oscillators.
If not given, drudeDamping is set to the value of langevinDamping.
- drudeBondLen
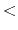 Drude oscillator bond length, beyond which to apply restraint (Å)
Drude oscillator bond length, beyond which to apply restraint (Å) 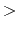
Acceptable Values: positive decimal
Description: An additional quartic restraining potential is applied to a Drude
oscillator if its length exceeds drudeBondLen.
The recommended value is 0.2 Å, fitted from QM calculations.
- drudeBondConst
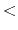 Drude oscillator restraining force constant
Drude oscillator restraining force constant 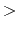
Acceptable Values: positive decimal
Description: If drudeBondConst is defined,
an additional quartic restraining potential is applied to a Drude
oscillator if its length exceeds drudeBondLen.
The recommended value is 40000, fitted from QM calculations.
- drudeNbTholeCut
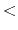 nonbonded Thole interaction radius (Å)
nonbonded Thole interaction radius (Å) 
Acceptable Values: positive decimal
Description: If drudeNbTholeCut is defined,
the screened Coulomb correction of Thole is also calculated
for non-excluded, nonbonded pairs of Drude oscillators that are
within this radius of interaction.
The recommended value is 5.0 Å for a high concentration (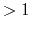 M)
ionic solution, or otherwise leave it 0.
M)
ionic solution, or otherwise leave it 0.




Next: MARTINI Residue-Based Coarse-Grain Forcefield
Up: Force Field Parameters
Previous: Water Models
Contents
Index
http://www.ks.uiuc.edu/Research/namd/