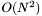 approach; for periodic systems it uses nearest image convention
approach; for periodic systems it uses nearest image convention
#include "mdapi/mdtypes.h"
Go to the source code of this file.
Classes | |
| struct | ForceParam_tag |
Used to configure Force_tag class. More... | |
| struct | ForceEnergy_tag |
| Container class returning potential energies. More... | |
| struct | ForceResult_tag |
| Container class returning atomic forces. More... | |
| struct | ForceCell_tag |
| (For internal use only.) More... | |
| struct | Force_tag |
| Force evaluation class. More... | |
Typedefs | |
| typedef ForceParam_tag | ForceParam |
Used to configure Force_tag class. | |
| typedef ForceEnergy_tag | ForceEnergy |
| Container class returning potential energies. | |
| typedef ForceResult_tag | ForceResult |
| Container class returning atomic forces. | |
| typedef ForceCell_tag | ForceCell |
| (For internal use only.) | |
| typedef Force_tag | Force |
| Force evaluation class. | |
Enumerations | |
| enum | ForceFlags_tag { FORCE_BOND = 0x000001, FORCE_ANGLE = 0x000002, FORCE_DIHED = 0x000004, FORCE_IMPR = 0x000008, FORCE_ELEC = 0x000010, FORCE_VDW = 0x000020, FORCE_BONDED = FORCE_BOND | FORCE_ANGLE | FORCE_DIHED | FORCE_IMPR, FORCE_NONBONDED = FORCE_ELEC | FORCE_VDW, FORCE_ALL = FORCE_BONDED | FORCE_NONBONDED, FORCE_ELEC_DIRECT = 0x000040, FORCE_VDW_DIRECT = 0x000080, FORCE_DIRECT = FORCE_ELEC_DIRECT | FORCE_VDW_DIRECT, FORCE_MASK_TYPE = 0x0000FF, FORCE_NONPERIODIC = 0x000000, FORCE_X_PERIODIC = 0x000100, FORCE_Y_PERIODIC = 0x000200, FORCE_Z_PERIODIC = 0x000400, FORCE_PERIODIC = FORCE_X_PERIODIC | FORCE_Y_PERIODIC | FORCE_Z_PERIODIC, FORCE_MASK_PERIOD = 0x000F00, FORCE_SPHERE = 0x001000, FORCE_X_CYLINDER = 0x002000, FORCE_Y_CYLINDER = 0x004000, FORCE_Z_CYLINDER = 0x008000, FORCE_MASK_BC = 0x00F000, FORCE_EXCL_NONE = 0x000000, FORCE_EXCL_12 = 0x010000, FORCE_EXCL_13 = 0x020000, FORCE_EXCL_14 = 0x030000, FORCE_EXCL_SCAL14 = 0x040000, FORCE_MASK_EXCL = 0x0F0000, FORCE_SMOOTH = 0x100000, FORCE_SWITCH = 0x200000, FORCE_CONTINUOUS = FORCE_SMOOTH | FORCE_SWITCH, FORCE_ELEC_EXCL = 0x400000, FORCE_VDW_EXCL = 0x800000, FORCE_EXCL = FORCE_ELEC_EXCL | FORCE_VDW_EXCL, FORCE_MASK_CUTOFF = 0xF00000, FORCE_MD_VACUUM = FORCE_ALL | FORCE_EXCL_SCAL14 | FORCE_CONTINUOUS, FORCE_MD_CELL = FORCE_MD_VACUUM | FORCE_PERIODIC } |
| Flags to indicate which parts of the force field are to be evaluated. More... | |
| enum | { FORCE_FAIL = -1 } |
Functions | |
| int | force_init (Force *) |
| Constructor. | |
| int | force_setup (Force *, ForceParam *fp, ForceEnergy *fe, ForceResult *fr) |
Setup the Force_tag object. | |
| int | force_compute (Force *, const MD_Dvec *pos, MD_Dvec *wrap) |
| Perform force evaluation. | |
| void | force_done (Force *) |
| Destructor. | |
| int32 ** | force_excl_list (const Force *) |
| Return exclusion lists. | |
| int32 ** | force_scaled14_list (const Force *) |
| Return scaled 1-4 exclusion lists. | |
| int | force_setup_bonded (Force *) |
| int | force_setup_nonbonded (Force *) |
| int | force_setup_nonbonded_cells (Force *) |
| int | force_setup_boundary (Force *) |
| int | force_compute_bonded (Force *, const MD_Dvec *pos) |
| int | force_compute_nonbonded (Force *, const MD_Dvec *pos, MD_Dvec *wrap) |
| int | force_compute_boundary (Force *, const MD_Dvec *pos, const MD_Dvec *wrap) |
Force_tag class evaluates the basic force field for biomolecules. Boundary conditions may be nonperiodic or orthogonal periodic (or semi-periodic) cells. Nonbonded interactions are treated using a cutoff distance. Helper class ForceParam_tag is used for initialization. Two helper classes ForceEnergy_tag and ForceResult_tag provide computed energies and forces, respectively.The force evaluation can compute subsets from the following:
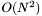 approach; for periodic systems it uses nearest image convention
approach; for periodic systems it uses nearest image conventionForce_tag class is used with three helper classes: ForceParam_tag is configured at the beginning of the simulation to describe which aspects of the force field are to be evaluated, along with force field parameter and molecule topology information ForceEnergy_tag returns the total potential energy along with parts of the potential energy (e.g. bond, angle, van der Waals) ForceResult_tag returns the total forces for each atom of the system and optionally the separated forces for each atom (e.g. bond, angle, van der Waals)force_compute() below) is always: pos[i] + wrap[i]
The units are as outlined by the MDAPI documentation:
Caveats:
Force_tag class, so this does not provide good support for constant pressure simulations.
|
|
Force evaluation class. Members should be treated as private. |
|
|
Container class returning potential energies.
User-supplied container class that returns potential energies from |
|
|
Used to configure
User sets the internal fields of this class in order to configure the evaluation performed by the
The user provides and retains ownership of memory buffer space for all arrays. The force field parameter and topology arrays all follow the MDAPI guidelines as defined by
The parameter The boundary restraint and nonbonded parameters follow the NAMD conventions, as described in its documentation.
The |
|
|
Container class returning atomic forces.
User-supplied container class that returns atomic forces from
The |
|
|
|
|
|
Flags to indicate which parts of the force field are to be evaluated.
Flags should be bitwise ORed together, used to initialize There are some values that collect commonly used flags:
If you want to use an external electrostatic library, you can use the FORCE_ALL | FORCE_EXCL_SCAL14 | FORCE_ELEC_EXCL | FORCE_SWITCH
|
|
||||||||||||||||
|
Perform force evaluation.
The convention here is that
|
|
|
Destructor.
Frees memory allocations made internally during |
|
|
Return exclusion lists.
This returns the exclusion lists created during |
|
|
Constructor.
Initializes a
|
|
|
Return scaled 1-4 exclusion lists.
The same format as returned by |
|
||||||||||||||||||||
|
Setup the
force_compute(). The user should not change these objects, particularly not the ForceParam_tag object, after calling this routine.
|
 1.4.4
1.4.4