From: Dhiraj Srivastava (dhirajks_at_gmail.com)
Date: Tue Oct 04 2016 - 16:07:43 CDT
Hi
I am trying to do MD simulation on a protein with and without ligand.
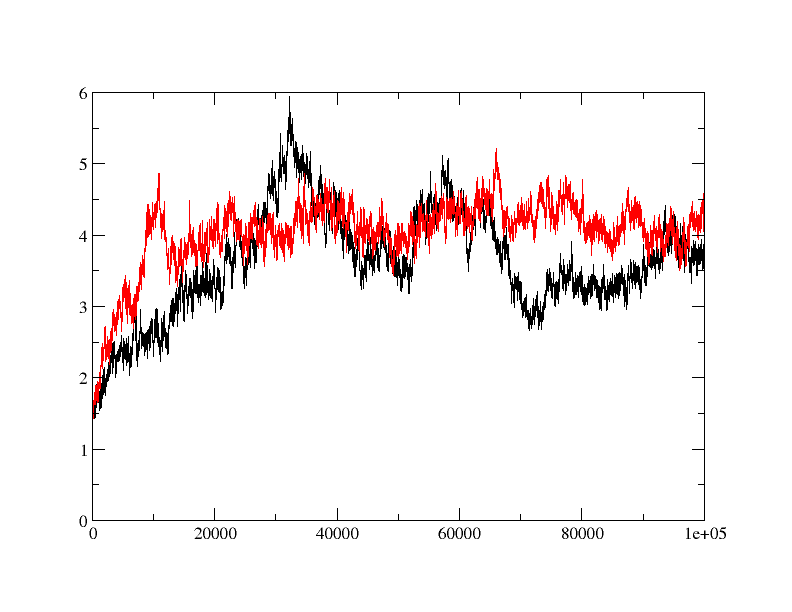
when I did rmsd plot, I found that apo protein is behaving fine (red) but
ligand bound form (black) is taking relatively longer time to equilibrate
and showing quite a bit of fluctuation in rmsd. is the fluctuation in rmsd
value for ligand bound protein is acceptable or is there anything wrong?
How can I fix it?
thanks
Dhiraj
[image: Inline image 3]

This archive was generated by hypermail 2.1.6 : Sun Dec 31 2017 - 23:20:44 CST