



Next: Moving Constraints
Up: User Defined Forces
Previous: External Electric Field
Contents
Index
NAMD provides the ability to specify grids describing a potential in the simulation
space. Each atom is affected by the potential based on its charge and its position,
using the potential function interpolated from the specified grid(s). Energy due to the
grid-defined field will be reported in the MISC column of the output, unless a scaling
factor not proportional to (1,1,1) is used.
NAMD allows the definition of multiple grids, each with a separate set of defining
parameters. This is specified using a tag field in each of the mgridforceXXX commands.
The tag is an alphanumeric string without spaces which identifies to which grid the
specified field applies.
The grid file format is a subset of the DataExplorer DX file format, as shown below:
# Lines at the beginning of the file starting with a # symbol
# are ignored as comments
# Variables (replaced by numbers in an actual file):
# xn, yn, and zn are the number of data points along each dimension;
# xorg, yorg, and zorg is the origin of the grid, in angstroms;
# x[1-3]del, y[1-3]del, and z[1-3]del are the basis vectors which transform
# grid indices to coordinates in angstroms:
# x(i,j,k) = xorg + i * x1del + j * y1del + k * z1del
# y(i,j,k) = yorg + i * x2del + j * y2del + k * z2del
# z(i,j,k) = zorg + i * x3del + j * y3del + k * z3del
#
# Grid data follows, with three values per line, ordered z fast, y medium,
# and x slow. Exactly xn*yn*zn values should be given. Grid data is then
# terminated with a field object.
#
# Note: Other features of the DX file format are not handled by this code
#
object 1 class gridpositions counts xn yn zn
origin xorg yorg zorg
delta x1del y1del z1del
delta x2del y2del z2del
delta x3del y3del z3del
object 2 class gridconnections counts xn yn zn
object 3 class array type double rank 0 items [ xn*yn*zn ] data follows
f1 f2 f3
f4 f5 f6
.
.
.
object 4 class field
component "positions" value 1
component "connections" value 2
component "data" value 3
Each dimension of the grid may be specified as continuous or not. If the grid is not continuous in a particular dimension, the potential grid is padded with one border slices on each non-continuous face of the grid, and border grid values are computed so that the force felt by an atom outside the grid goes to zero. If the grid is continuous along a particular dimension, atoms outside the grid are affected by a potential that is interpolated from the grid and its corresponding periodic image along that dimension.
To calculate the force on an atom due to the grid, the atom's coordinates are transformed according to the current basis vectors of the simulation box to a coordinate frame that is centered at the center of the specified grid. Note that the size and spatial coordinates of the grid remain fixed, and are not scaled as the size of the simulation box fluctuates. For atoms within the grid, the force is computed by analytically determining the gradient of the tricubic polynomial used to interpolate the potential from surrounding grid values. For atoms outside the grid, the state of the mgridforcecont[1,2,3] determine whether the force is zero, or computed from the images of the grid as described above. Note that if the grid is ever larger than the periodic box, it is truncated at the edge of that box. The consequence of this is that the computed potential will not vary smoothly at the edges, introducing numerical instability.
NAMD also supports non-uniform grids, allowing regions of a grid to be defined at higher resolution.
Non-uniform grids are structured hierarchically, with a single maingrid which has one or more subgrids.
Each subgrid spans a number of maingrid cells in each of the three dimensions, and effectively redefines the data in that region.
The subgrids are usually defined at higher resolution, with the restriction that the number of cells along each dimension is an integral number of the original number in the maingrid.
Note that the maingrid still has data points in regions where subgrids are defined, and that, on the boundary of a subgrid, they must agree with the values in the subgrid.
Subgrids, in turn, may have subgrids of their own, which may have subgrids of their own, etc.
A non-uniform grid file takes the form of a special comment block followed by multiple normal grid definitions.
The special comment block defines the grid hierarchy, and consists of comments beginning with # namdnugrid.
An example follows:
# namdnugrid version 1.0
# namdnugrid maingrid subgrids count 2
# namdnugrid subgrid 1 generation 1 min x1 y1 z1 max x2 y2 z2 subgrids count 2
# namdnugrid subgrid 2 generation 2 min x3 y3 z3 max x4 y4 z4 subgrids count 0
# namdnugrid subgrid 3 generation 2 min x5 y5 z5 max x6 y6 z6 subgrids count 0
# namdnugrid subgrid 4 generation 1 min x7 y7 z7 max x8 y8 z8 subgrids count 0
The maingrid is described by the number of subgrids.
Subgrids are additionally described by a subgrid number; a generation number, which should be one higher than the generation of its supergrid; and min and max attributes, which describe the location of the subgrid within its supergrid.
In this example, the maingrid has two subgrids, subgrid 1 and subgrid 4, labeled generation 1.
The first of these subgrids has two subgrids of its own (generation 2).
Notice that subgrids are described immediately after their supergrid.
The min and max attributes are given in units of grid cells of the supergrid.
For example, a subgrid with min 0 0 0 max 1 1 1 would redefine 8 cells of its supergrid, the space between gridpoints (0, 0, 0) and (2, 2, 2) in grid coordinates.
Following the comment block, the maingrid and subgrids are defined in the format described above, in the same order as the comment block.
The following parameters describe the grid-based potentials.
- mgridforce
 apply grid forces?
apply grid forces? 
Acceptable Values: yes or no
Default Value: no
Description: Specifies whether or not any grid forces are being applied.
- mgridforcefile
 tag
tag 
 PDB file specifying force multipliers and charges for each atomd
PDB file specifying force multipliers and charges for each atomd 
Acceptable Values: UNIX file name
Description: The force on each atom is scaled by the corresponding value in this PDB file. By setting the force multiplier to zero for an atom, it will not be affected by the grid force.
- mgridforcecol
 tag
tag 
 column of PDB from which to read force multipliers
column of PDB from which to read force multipliers 
Acceptable Values: X, Y, Z, O, or B
Default Value: B
Description: Which column in the PDB file specified by mgridforcefile contains the scaling factor
- mgridforcechargecol
 tag
tag 
 column of PDB from which to read atom charges
column of PDB from which to read atom charges 
Acceptable Values: X, Y, Z, O, or B
Default Value: Atom charge used for electrostatics.
Description: Which column in the PDB file specified by mgridforcefile contains the atom charge. By default, the charge value specified for the short-range Columb interactions are also used for the grid force. Both mgridforcecol and mgridforceqcol can be specified, in which case the apparent charge of the atom will be the product of the two values.
- mgridforcepotfile
 tag
tag 
 grid potential file name
grid potential file name 
Acceptable Values: UNIX file name
Description: File specifying the grid size, coordinates, and potential values.
- mgridforcevolts
 tag
tag 
 grid potential units in eV/charge
grid potential units in eV/charge 
Acceptable Values: yes or no
Default Value: no
Description: If set, the grid potential values are expressed in eV. Otherwise, values are
in
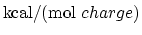
- mgridforcescale
 tag
tag 
 scale factor for grid potential
scale factor for grid potential 
Acceptable Values: Vector of decimals scale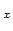 scale
scale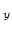 scale
scale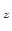
Default Value: 1 1 1
Description: Scale factor applied to the grid potential values
- mgridforcecont1
 tag
tag 
 Is grid continuous in the direction of the first basis vector
Is grid continuous in the direction of the first basis vector 
Acceptable Values: yes or no
Default Value: no
Description: By specifying that the grid is continuous in a direction, atoms outside of the grid will be affected by a force determined by interpolating based on the values at the edge of the grid with the values of the corresponding edge of the periodic image of the grid. The current size of the simulation box is taken into account, so that as the simulation box size fluctuates, the force on an atom outside of the grid varies continuously until it re-enters the opposite edge of the grid. If the grid is not continuous in this direction, the interpolated force on atoms near the edge of the grid is calculated so that it continuously approaches zero as an atom approaches the edge of the grid.
- mgridforcecont2
 tag
tag 
 Is grid continuous in the direction of the second basis vector
Is grid continuous in the direction of the second basis vector 
Acceptable Values: yes or no
Default Value: no
Description: Operates the same as mgridforcecont1 except applies in the direction of the second basis vector
- mgridforcecont3
 tag
tag 
 Is grid continuous in the direction of the third basis vector
Is grid continuous in the direction of the third basis vector 
Acceptable Values: yes or no
Default Value: no
Description: Operates the same as mgridforcecont1 except applies in the direction of the third basis vector
- mgridforcevoff
 tag
tag 
 Offset periodic images of the grid by specified amounts
Offset periodic images of the grid by specified amounts 
Acceptable Values: vector of decimals (x y z)
Default Value: (0 0 0)
Description: If a continuous grid is used along a particular basis vector, it may be desirable to shift the potentials in the image to manipulate the potential outside the grid. For example, consider the case where the potential is a ramp in the 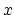 direction and the grid is defined for points
direction and the grid is defined for points 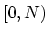 , with a potential
, with a potential 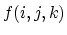 given by
given by
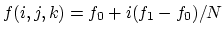 . By shifting the images of the grid, the potential can be transformed as illustrated in Fig. 5.
. By shifting the images of the grid, the potential can be transformed as illustrated in Fig. 5.
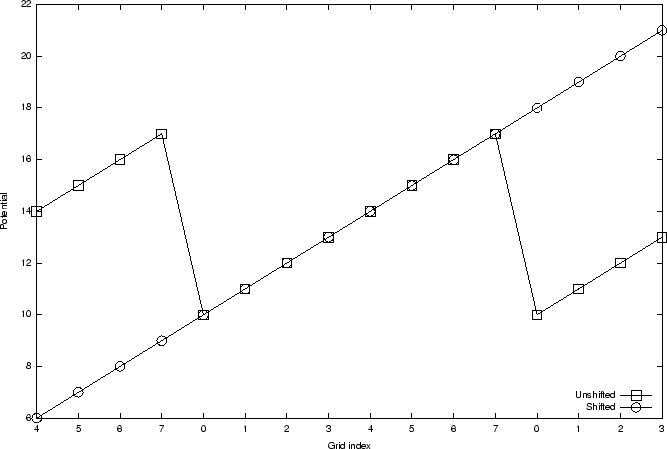
Figure 5:
Graph showing a slice of a ramp potential, with eight grid points along the axis, and a periodic cell size which just contains the grid. The Unshifted case shows how the pontential is not smooth when mgridforcevoff is not specified, or set to zero. The Shifted potential shows the grid that results when mgridfocevoff is set so that the wrapped potential is offset so that the potential has constant slope at the periodic boundaries.
 |
- mgridforcelite
 tag
tag 
 Is grid to use Gridforce Lite interpolation?
Is grid to use Gridforce Lite interpolation? 
Acceptable Values: yes or no
Default Value: no
Description: When Gridforce Lite is enabled, a faster but less accurate interpolation method is used to compute forces.
Specifically, rather than computing a tri-cubic interpolation of the potential, from which the force is then computed analytically, Gridforce Lite computes force as a linear interpolation.
This method also increases the memory required by Gridforce.
Note that Gridforce Lite is incompatible with use of the mgridforcecont[123] keywords and with non-uniform grids.




Next: Moving Constraints
Up: User Defined Forces
Previous: External Electric Field
Contents
Index
http://www.ks.uiuc.edu/Research/namd/
