|
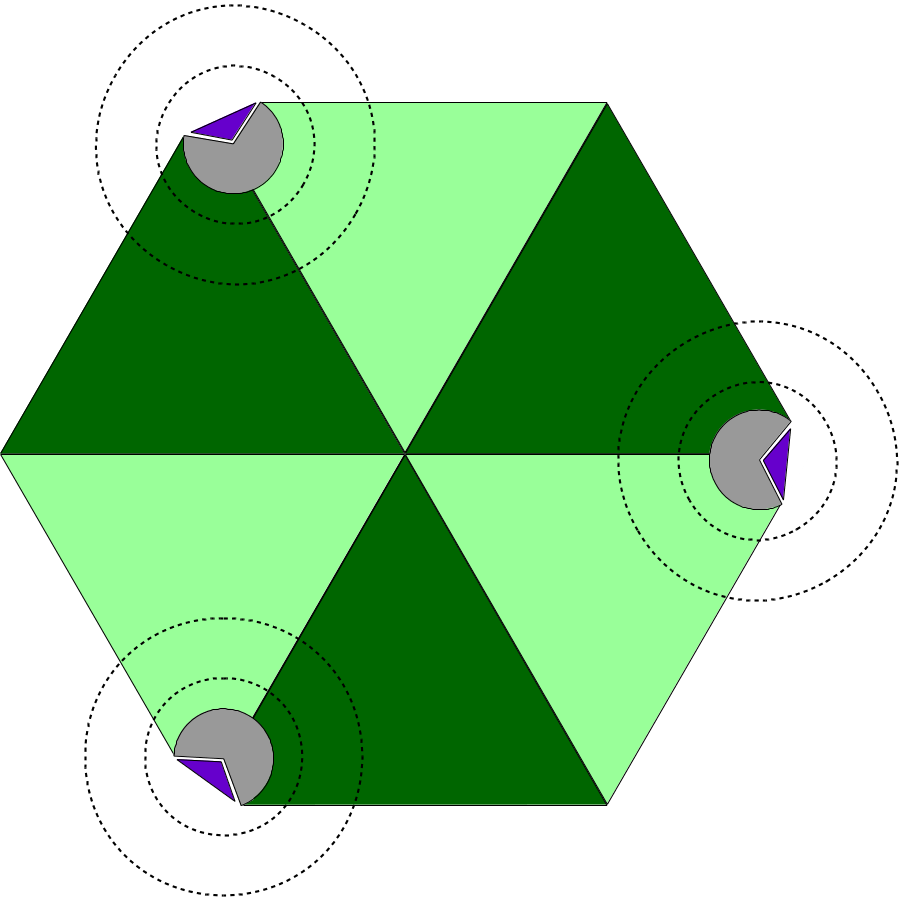
Aiming at large macromolecular simulations that could take advantage of localized QM resolution, NAMD allows the user to set up multiple independent QM regions in the same molecular system. For example, one could study a multimeric complex that contains several active sites and have all active sites be calculated with a chosen QC software simultaneously (Figure 19). Each active site would be calculated independently of all others, by its own execution of the QC software, keeping the calculation cost low and without impacting the overall efficiency of the simulation, since all QM regions would be calculated in parallel.

|
Identifying the different QM regions and which atoms belong to each one of them can be simply accomplished in the input PDB file, or in a dedicated PDB file (keyword ``qmParamPDB"). Since each region can contain different sets of molecules, their charges and multiplicities are indicated separately (see keywords ``qmCharge" and ``qmMult").
For simulations of large systems that are distributed across several computer nodes, one can control how many independent QM regions are calculated in each node. This would prevent large simulations from running out of memory if two or more large QM regions are placed in the same node (see keyword ``qmSimsPerNode").