- Choosing a function
- distance: center-of-mass distance between two groups.
- distanceZ: projection of a distance vector on an axis.
- distanceXY: modulus of the projection of a distance vector on a plane.
- distanceVec: distance vector between two groups.
- distanceDir: distance unit vector between two groups.
- distanceInv: mean distance between two groups of atoms.
- distancePairs: set of pairwise distances between two groups.
- cartesian: vector of atomic Cartesian coordinates.
- angle: angle between three groups.
- dipoleAngle: angle between two groups and dipole of a third group.
- dihedral: torsional angle between four groups.
- polarTheta: polar angle in spherical coordinates.
- polarPhi: azimuthal angle in spherical coordinates.
- coordNum: coordination number between two groups.
- selfCoordNum: coordination number between atoms within a group.
- hBond: hydrogen bond between two atoms.
- rmsd: root mean square displacement (RMSD) from reference positions.
- Advanced usage of the rmsd component.
- Path collective variables
- eigenvector: projection of the atomic coordinates on a vector.
- gyration: radius of gyration of a group of atoms.
- inertia: total moment of inertia of a group of atoms.
- inertiaZ: total moment of inertia of a group of atoms around a chosen axis.
- orientation: orientation from reference coordinates.
- orientationAngle: angle of rotation from reference coordinates.
- orientationProj: cosine of the angle of rotation from reference coordinates.
- spinAngle: angle of rotation around a given axis.
- tilt: cosine of the rotation orthogonal to a given axis.
- alpha:
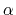 -helix content of a protein segment.
-helix content of a protein segment.
- dihedralPC: protein dihedral pricipal component
- Shared keywords for all components
- Periodic components
- Non-scalar components
- Linear and polynomial combinations of components
- Custom functions
- Scripted functions
- Defining grid parameters
- Trajectory output
- Extended Lagrangian
- Backward-compatibility
- Statistical analysis
- Atom selection keywords
- Moving frame of reference.
- Treatment of periodic boundary conditions.
- Performance of a Colvars calculation based on group size.
- Adaptive Biasing Force
- Extended-system Adaptive Biasing Force (eABF)
- Metadynamics
- Harmonic restraints
- Computing the work of a changing restraint
- Harmonic wall restraints
- Linear restraints
- Adaptive Linear Bias/Experiment Directed Simulation
- Multidimensional histograms
- Probability distribution-restraints
- Defining scripted biases
- Performance of scripted biases