Animations of ATP Synthase MD Simulations
The following movies illustrate the MD simulations reported in the article.
The thumbnail images are linked to the movies in MPEG 1 format; use links
under the images to view movies in Quicktime format.
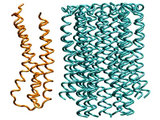
Fitting subunit a to the c10 oligomer
Subunit a undergoes 10,000 conjugate gradient minimization steps
followed by equilibration in vacuum at 4K for 130ps. All backbone atoms of
all subunits c are restrained. The equilibration at the low temperature
assures that no major conformation change could occur, while resulting in
a well adjusted interface between the a subunit and
the c10 oligomer.
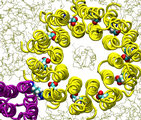
Forced rotation of the c10 oligomer
Oligomer c10 is forced to rotate relative to subunit
a and surrounding lipids. Steering forces are applied to all backbone
atoms of the c10 oligomer. The total torque exerted
on c10 is 2,600 Kcal/mol, and the total simulation time
is 1ns. In this simulation, the proton binding sites in all c
subunits are protonated. The rotation angle of the c10oligomer
is calculated by averaging the rotation angles for each csubunit,
which were themselves computed using the subunit center of mass positions
at time t and at the beginning of the MD simulations.
Forced rotation of the c2 helix
Steering forces are applied to the backbone atoms of residues 47 through
79 (cTMH2), while the backbone atoms of cTMH1 are restrained.
The average torque exerted on the backbone of cTMH2 is 300Kcal/mol
in simulation a), and 60 kcal/mol in simulation b). The total
simulation times are 100ps and 6ns, respectively. Residues 65, 61, 57 and
53 are shown in the van der Waals representation. In movie b), motion
of the protein was averaged over a 100ps window.
Deprotonation of a single cAsp61 residue blocks rotation of the
c10 oligomer
With only one cAsp61 at the interface deprotonated, a rapid (within
about 10ps) formation of a salt bridge with aArg210 is observed. The
salt bridge ties aTMH4 to the c10 oligomer, tearing aTMH4
off the other TMH's in the a subunit as the forced rotation of the
c10 oligomer continues. In a similar simulation, when all backbone
atoms of the a subunit are restrained, the outer TMH of the c
subunit quickly unwinds.
Salt bridge transfer between two deprotonated cAsp61 residues
At the outset cAsp61 of c2R was deprotonated forming a
salt bridge with cArg210. Helix c2L was rotated counterclockwise
by 180o in a 1 ns simulation (the same result could be achieved
by a 180o rotation in the opposite direction). When cAsp61
of c2L approached the terminal residue aSer206 of the cytoplasmic
channel, it was deprotonated, mimicking proton release to the cytoplasm.
At this point, a complex of three charged residues formed, dramatically reducing
the dissociation energy of the salt bridge between aArg210 and cAsp61
and, thereby, making it possible to transfer the cAsp61-aArg210
salt bridge from one c subunit to the other. At this point, cAsp61
at c2R, which formed a hydrogen bond with the terminal residue
aAsn214 of the periplasmic channel, was protonated, mimicking proton
intake from the periplasm; the c10 oligomer rotated counterclockwise
(synthesis direction) by about 36o, and helix c2L rotated
clockwise by 180o. For technical reasons, the oligomer and the
outer TMH were rotated in steps one after the other rather than simultaneously.
The salt bridge between aArg210 and cAsp61 at c2L
stayed intact and no significant distortions of the structure were observed,
i.e., the system returned to the starting conformation, with the c10
oligomer advanced by 36o.
Design by Ilya Balabin
Last updated May 29, 2003.