Highlights of our Work
2024 | 2023 | 2022 | 2021 | 2020 | 2019 | 2018 | 2017 | 2016 | 2015 | 2014 | 2013 | 2012 | 2011 | 2010 | 2009 | 2008 | 2007 | 2006 | 2005 | 2004 | 2003 | 2002 | 2001
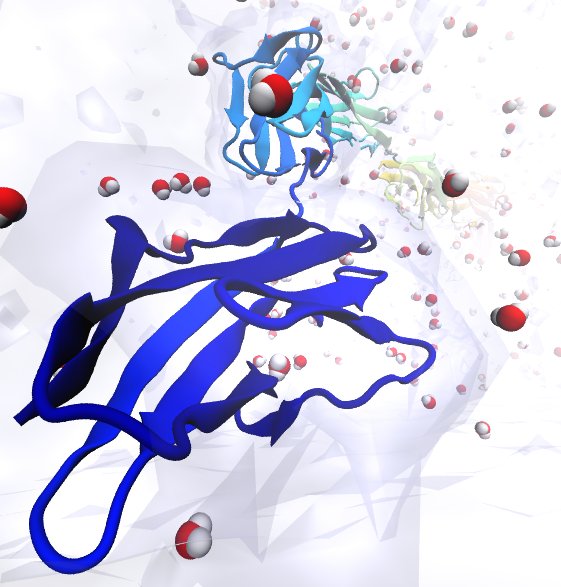
image size:
60.3KB
made with VMD
A cell that is alive can be recognized typically under a microscope through cell motion, streaming of inner cell fluids, and many other mechanical cell activities. These activities are the result of mechanical forces that arise in the cell, but even though the effect of the forces on cells are tremendous, the magnitude of the forces when measured by physical instruments, are extremely small, 1/1000000000000 of the force felt when lifting a Kilogramm. Sensing such small force requires extremely high precision in research techniques, and, for more than a decade, computational biologists, using NAMD, have exploited the all-atom resolution of molecular dynamics modeling to explain very successfully the molecular level effect of small forces in cells (for example, see our previous highlights on muscle protein, blood clot protein, and other successful applications). But simulations, too, are challenged by the small magnitude of cellular forces, mainly because precise simulation requires extremely extensive simulations, that cover a duration as close as possible to the biological time scale. Advances in computational biology have now permitted in the case of a the muscle protein titin a simulation of unprecedented accuracy that resolved clearly the relationship between molecular structure of titin and its ability to sustain the forces that arise in muscle function. For this purpose a key element of titin was stretched at a velocity slow enough that hydrodynamic drag, that came about as an unwanted byproduct of earlier simulations, was negligible compared to titin's intrinsic force bearing properties. The new simulations opened an unveiled view on muscle elasticity. More information can be found on our titin website, and in a recent review .



